| Issue |
Parasite
Volume 32, 2025
|
|
|---|---|---|
| Article Number | 61 | |
| Number of page(s) | 12 | |
| DOI | https://doi.org/10.1051/parasite/2025053 | |
| Published online | 25 September 2025 | |
Research Article
Seasonal exchange of microsporidian parasites between native and non-native pet-traded freshwater crustaceans: Is parasite spillover favored over spillback?
Échanges saisonniers de microsporidies parasites entre crustacés d’eau douce indigènes et crustacés non indigènes commercialisés comme animaux de compagnie : le débordement parasitaire est-il favorisé par rapport à la retransmission ?
1
Department of Aquatic Ecology, University of Duisburg-Essen, Universitätsstr. 5, 45141 Essen, Germany
2
Centre for Water and Environmental Research, University of Duisburg-Essen, Universitätsstr. 2, 45141 Essen, Germany
3
Université de Montpellier, Place Eugène Bataillon, 34095 Montpellier, France
4
Reserva: The Youth Land Trust, P.O. Box 57277, Washington, DC 20037, USA
5
Association Guibétois, 211 Chemin du triol, 34343 Viols-le-Fort, France
6
Université de Toulouse (Paul Sabatier), 118 Route de Narbonne, 31400 Toulouse, France
7
Research Center One Health Ruhr of the University Alliance Ruhr, University of Duisburg-Essen, Universitätsstr. 5, 45141 Essen, Germany.
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
25
June
2025
Accepted:
25
August
2025
Abstract
The introduction of non-native pet-traded species poses potential threats to global biodiversity, particularly in freshwater ecosystems. This study investigated the seasonal dynamics of microsporidian infections in an established feral population of cherry shrimp (Neocaridina davidi) and the coexisting populations of crustaceans, comprising both native and non-native species, inhabiting the thermal waters of the Fontcaude Park and the nearby Mosson River in southern France. Our aim was to assess the potential occurrence of spillover and/or spillback events between N. davidi and co-occurring crustaceans, as well as the influence of seasonal dynamics on these interactions. The prevalence and diversity of microsporidian parasites exhibited strong seasonal variations. Although parasites associated with the pet trade were not detected, we highlight the acquisition of native parasites by feral N. davidi, which seems to be a suitable alternative host for native host-generalist microsporidians. Our findings indicate that all prerogatives for spillback events to occur are met. Feral N. davidi may establish and survive year-round in European rivers with natural thermal regimes. Thus, human-mediated introductions can potentially alter parasite transmission dynamics in these ecosystems.
Résumé
L’introduction d’espèces non indigènes, commercialisées comme animaux de compagnie, constitue une menace potentielle pour la biodiversité mondiale, en particulier dans les écosystèmes d’eau douce. Cette étude examine la dynamique saisonnière des infections microsporidiennes au sein d’une population férale établie de crevettes de David (Neocaridina davidi) et des populations coexistantes de crustacés, indigènes et non indigènes, vivant dans les eaux thermales du parc de Fontcaude et de la Mosson, dans le sud de la France. Notre objectif était d’évaluer la survenue potentielle de débordement parasitaire et/ou de retransmission parasitaire entre N. davidi et les crustacés coexistants, ainsi que l’influence de la dynamique saisonnière sur ces interactions. La prévalence et la diversité des microsporidies parasites ont montré de fortes variations saisonnières. Bien que les parasites associés au commerce des animaux de compagnie n’aient pas été détectés, nous soulignons l’acquisition de parasites indigènes par N. davidi de la population férale, qui semble être un hôte alternatif approprié pour les microsporidies généralistes indigènes. Nos résultats indiquent que toutes les conditions sont réunies pour que des événements de retransmission se produisent. Neocaridina davidi pourrait s’établir et survivre toute l’année dans les rivières européennes à régimes thermiques naturels. Ainsi, les introductions d’origine humaine peuvent potentiellement modifier la dynamique de transmission des parasites dans ces écosystèmes.
Key words: DNA barcoding / Freshwater ecosystems / Invasive non-native species / Microsporidia / Neocaridina davidi / Parasite transmission
Edited by: Jean-Lou Justine
© S. Prati et al., published by EDP Sciences, 2025
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Introduction
The introduction of non-native pet-traded species is a growing phenomenon and a significant threat to global biodiversity [11, 15, 49, 58]. Once introduced, such species might be able to establish self-sustaining populations and eventually spread to other areas, becoming invasive, often asserting significant impacts on the local ecosystems [11, 15]. Freshwater ecosystems are especially affected by invasive species, which often lead to substantial biodiversity loss and major changes in energy flows and the physical environment [7, 15, 16, 21]. Direct impacts asserted by invasive species on receiving ecosystems typically involve biotic interactions such as competition, predation, hybridization, and pathogen transmission [7]. In contrast, indirect impacts often include changes in habitat structure, water clarity, organic matter, and nutrient concentrations [7]. Due to a complex interplay of a wide range of processes, including pathogen transmission and possible asymmetry in direct and indirect impact trajectories, the introduction of non-native pet-traded species with a known invasion history may lead to unpredictable outcomes.
The rise of the pet trade as a major pathway for the introduction of non-native species and their associated symbionts may enhance pathogen transmission between non-native and native species. This risk is exacerbated by insufficient biosecurity measures and potential immunosuppression of farmed pet animals, notably due to inbreeding and the relatively high density of individuals in aquaculture facilities, which can lead to higher pathogen prevalence compared to wild populations [38, 40, 49, 72]. For instance, ornamental cyprinids, such as Carassius auratus (Linnaeus, 1758) and Cyprinus carpio Linnaeus, 1758, are believed to be responsible for the spread of the anchor worm Lernaea cyprinacea Linnaeus, 1758, a crustacean parasite causing extensive hemorrhage and ulcerations in native Australian fish species [20]. Moreover, the introduction of non-native crayfish species of North American origin, including those linked to the pet trade, has led to severe declines and even extinctions of numerous native European crayfish species. A dire situation primarily caused by interspecific competition and the lack of immune defenses in native species against the crayfish plague, a deadly disease vectored by North American crayfish [43, 62]. As pathogens often exhibit broader environmental tolerances than their hosts [13], the introduction of non-native pet-traded species may facilitate the spread of pathogens beyond established climatic boundaries. Most pet-traded species originate from tropical or subtropical regions. They may struggle to establish themselves in colder climates, such as those found at higher latitudes (e.g., Europe, Northeast Asia, or North America). Still, co-introduced pathogens might occasionally persist in novel ecosystems, even after their original host has disappeared [8, 58].
The “enemy release” theory suggests that non-native species partially lose their associated pathogens when introduced into new environments [65]. Pathogens may be lost due to handling, the subsampling effect (only uninfected individuals reach the transport stage), or during transport [11, 65]. After introduction, pathogens may fail to establish themselves because they either die shortly after, lack the ability to reproduce or have a reproduction rate too low to create a self-sustaining population [11]. Establishment can be especially challenging for non-native pathogens with complex life cycles, as they often depend on multiple specific hosts that may be absent in the receiving ecosystem [11, 65]. However, parasites and other pathogens with simple life cycles (i.e., requiring only one host), particularly host-generalists, are more likely to persist in novel environments [53]. If the remaining pathogens are host-specialists, they may be constrained to the introduced non-native species and possibly a small number of closely related native host species; however, if they are host-generalists, spillover events to susceptible native species may occur. If multiple pathogens are introduced alongside an invasive host, a potential invasional meltdown, though rare, may occur [11]. Conversely, non-native species might also acquire local parasites due to a lack of immune defenses and/or close relatedness with native species, amplifying (spillback) or diluting (dilution) their prevalence in native populations [11, 29, 61, 63]. Thereafter, parasite transmission between non-native and native species is particularly relevant when dealing with potential pathogen reservoirs such as commercially successful freshwater pets. The latter, often immunosuppressed, may host generalist parasites with simple life cycles, and are more likely to be released into natural environments through aquarium dumping practices, enhancing the risk of pathogen spillover [17, 24, 43].
Popular pet-traded freshwater crustaceans such as crayfish (Astacidae) and shrimp (Atyidae) are known carriers of a wide range of pathogens and have established viable feral populations worldwide [4, 23, 40, 52]. One notable example is the commercially successful cherry shrimp Neocaridina davidi (Bouvier, 1904) (syn. N. denticulata sinensis and N. heteropoda), a freshwater species of Southeast Asian origin known for its high fecundity and environmental plasticity [52]. Feral self-sustaining N. davidi populations are currently found in Canada, Germany, France (metropolitan area and in the overseas territory of La Réunion), Hungary, Israel, Japan, Poland, Slovakia, and the USA (Hawaii) [24, 51, 52]. While invasive at lower latitudes, in continental Europe, N. davidi is thought to be limited to thermal waters, but recent evidence shows that, aided by a warming climate, it is expanding into colder waters [52]. Known ecological impacts of feral N. davidi include the replacement of native species with similar ecological niches [47], meiofaunal assemblage alterations [69], changes in the leaf-litter breakdown and thus energy flows in invaded areas [57], and possible parasite exchange with native and/or non-native species [52].
Neocaridina davidi hosts a wide range of commensals and parasites [3, 34, 40, 46, 52], some of which have already been co-introduced outside their native range [26, 39, 45, 52, 56]. Among them is the ecologically and economically relevant microsporidian parasite Ecytonucleospora hepatopenaei (Tourtip et al., 2009) Wang et al., 2023 (syn: Enterocytozoon hepatopenaei), which has been detected in pet-traded and feral N. davidi populations across Europe [52, 56]. This host-generalist parasite with a simple life cycle infects many freshwater, brackish, and marine invertebrates, often causing substantial economic losses in the shrimp aquaculture industry [10, 25, 27, 32, 44]. Recent evidence suggests that microsporidian parasites may be shared between feral N. davidi and the invasive Red Swamp crayfish (Procambarus clarkii (Girard, 1852) [52]. However, it is unclear whether spillover or spillback events may occur between feral N. davidi and native species in invaded ecosystems. An attempt to experimentally infect native amphipods and isopods using N. davidi tissues containing E. hepatopenaei spores was not successful [52]. Still, feral N. davidi can host microsporidian parasites phylogenetically close to those found in insects such as coleopterans and lepidopterans, suggesting that spillback events might occur [51]. Yet, parasitological investigations comparing the parasite communities of feral N. davidi with those of native species sharing the same habitat are scarce and consist only of snapshots, thereby not accounting for potential seasonal dynamics [52]. As with other parasites, microsporidian infections can display pronounced seasonality [50], potentially leading to undetected spillover or spillback events in studies with suboptimal sampling periods.
Hence, we decided to investigate the microsporidian fauna of a recently discovered feral population of N. davidi inhabiting the warm waters of the Fontcaude thermal park of Juvignac (Southern metropolitan France) since at least 2021 [24]. Its water hosts both native species, such as isopods and amphipods, which are known hosts for a wide range of microsporidian parasites, many of which are host generalists [18, 53], as well as fish and crustacean species of pet-traded origin [24]. The thermal water originating from a spring feeds into a pond and then outflows in a ~ 70 m long channel, gradually losing heat before being redirected to the nearby Mosson River via an underground pipe and/or surface overflow channels in the downstream section [24]. These settings provide an ideal environment for microsporidian parasite exchanges between native and non-native species, and eventually, the spread of propagules in the Mosson River.
Profiting from this unique opportunity, the current study aimed to assess whether microsporidian spillover and/or spillback events may have occurred between N. davidi and other native and non-native crustaceans living in the park and the nearby Mosson River, and whether seasonality influenced these events. To this end, we employed DNA barcoding to molecularly characterize native and non-native crustaceans and their microsporidian parasites across different seasons. Additionally, pet-traded shrimps and their parasites were DNA-barcoded to facilitate direct comparison with feral individuals. We first hypothesized that microsporidian parasites found in pet-traded N. davidi would be detected in the feral population, as observed elsewhere in Europe [51, 52, 56]. Second, we hypothesized that spillback would be more favored than spillover events due to the higher diversity and prevalence of generalist parasites in native species compared to N. davidi and a limited subset of introduced animals, possibly stemming from uninfected pet-traded individuals. Last, we hypothesized that the prevalence and diversity of microsporidians in both native and non-native species would vary between seasons.
Material and methods
Sampling and processing
Between March and November 2024, N. davidi individuals and co-occurring amphipods, decapods, and isopods were collected using hand nets and baited bottles in the Fontcaude thermal park of Juvignac (Hérault, France; 43.627707, 3.811910) following the French fishing act and a prefectural authorization (id: DDTM34-2024-05-14915). At this site, warm water originating from a thermal spring-fed pond flows through a channel toward the Mosson River, gradually cooling along its course. This environment supports the coexistence of native and non-native species, including several taxa introduced via the pet trade, such as N. davidi and the guppy, Poecilia reticulata Peters, 1859 [24]. Neocaridina davidi was first recorded in the thermal park in December 2023 and has maintained a stable population since then. Its presence, however, is believed to date back at least to 2021 [24]. Individuals of N. davidi have been observed year-round in both the warmer and colder areas of the water body, suggesting that, if able to reach the nearby Mosson River, they might survive in it. However, their presence in the Mosson River had not been previously investigated; thus, in September and November 2024, we sampled a section of the river spanning 500 m upstream and downstream of the thermal park channel outlet, using baited bottles in accordance with the French fishing act. Unfortunately, most of the samples collected in the Mosson in September were later stolen, along with part of the equipment. Fifty-two individuals of N. davidi were also purchased from local pet shops and private sellers, as well as one of Europe’s largest and most popular online stores in Germany. All crustaceans employed in the study were euthanized using eugenol [9], and upon death, they were fixed in 96% ethanol for molecular analyses of both hosts and parasites. All hosts were measured and dissected at the Aquatic Ecology laboratory of the University of Duisburg-Essen (Germany). The processing of samples included visual screening for internal parasites, gut removal to prevent microsporidian contamination, and the dissection of a small portion of muscle and hepatopancreatic tissue for downstream molecular analyses. Epibionts were not analyzed as their conservation in ethanol-preserved specimens is often poor [52].
Host DNA extraction and amplification
We isolated DNA from hosts using a modified salt precipitation protocol [19]. Host molecular identification was performed using the universal eukaryotic primers LCO1490 (5′–GGTCAACAAATCATAAAGATATTGG–3′) and HCO2198 (5′–TAAACTTCAGGGTGACCAAAAAATCA–3′) [14], targeting the cytochrome oxidase I (COI) region. This primer pair has previously been used to amplify a wide range of crustaceans, including those analyzed in this study [52]. Additionally, we developed novel primers for N. davidi, targeting the mitochondrial 12S and 16S regions, based on published mitogenome sequences (NCBI GenBank accession numbers MN418055, NC_043865, and MK907783), to enhance the availability of alternative markers for future phylogenetic analyses. These were the following: 12SNeoF (5′–AAAGTGCGGGTTAAGATTGTGC–3′) and 12SNeoR (5′–TCCAGCACACCTACCTTGTTAC–3′), 16SNeoF (5′–TTGGCATCTCGAAGTGGAATGA–3′) and 16SNeoR. PCR reactions for hosts consisted of 20 μL assay with 10 μL of Dream-TaqTM Hot Start Green PCR Master Mix (Thermo Fisher Scientific, Waltham, MA, USA), 1.6 μL (5 μM) of each primer, 4.8 μL of nuclease-free water, and 2 μL of DNA template per reaction. PCR settings used for the primer pairs LCO1490–HCO2198 followed those used by [51], while those used for the newly developed primers targeting the 12S and 16S regions were as follows: initial denaturation for 2 min at 95 °C, followed by 30 cycles of 30 s denaturation at 95 °C, 40 s annealing at 59 °C and 90 s elongation at 72 °C, with a final elongation of 8 min at 72 °C.
Microsporidian DNA amplification
Microsporidian parasites were identified using the universal primers V1F (5′–CACCAGGTTGATTCTGCCTGAC–3′) [74] and 1342R (5′–ACGGGCGGTGTGTACAAAGAACAG–3′) [41], targeting the small subunit ribosomal RNA gene (SSU rRNA). PCR reactions for microsporidians consisted of 20 μL composed of 10 μL of 2× AccuStart II PCR ToughMix (Quantabio, Beverly, MA, USA), 1 μL of each primer (0.5 μM), 0.35 μL of 50× GelTrack Loading Dye (Quantabio), 6.65 μL MilliQ water, and 1 μL of DNA template. PCR settings used for the primer pairs V1F-1342R followed those used by [51].
Sequences and phylogenetic analyses
PCR products from hosts and microsporidians were sent unpurified to Microsynth Seqlab (Göttingen, Germany) for Sanger sequencing using LCO1490, 12SneoF, 16SneoF, and V1 primers, respectively. Raw sequences were quality-checked and edited using Geneious v2025.0.2 (GraphPad Software LLC d.b.a Geneious) and compared against GenBank records using BLASTN (https://blast.ncbi.nlm.nih.gov/). Host and parasite sequences were aligned using the MAFFT v7.490 algorithm with standard settings [28]. Maximum likelihood phylogenetic trees with bootstrap support values (1,000 replicates) were produced in IQ-Tree v2.4.0 [42] using the HKY + F + I substitution model for crustaceans and the TIM3 + F+I + R4 substitution model for microsporidians, both based on Bayesian Information Criterion scores. The respective outgroups were Neocaridina palmata (Shen, 1948) (MN701612) and Metchnikovella dogieli (Paskerova et al., 2016) (MT969020). The names and circumscriptions of N. davidi clades followed those proposed by Prati et al. [52], while those of microsporidian orders followed Bojko et al. [5]. Following the naming used in previous works [50–56], we used the artificial name “Microsporidium sp.” for all microsporidian isolates lacking a formal description. The only exception was for undescribed isolates belonging to the genus Nosema, for which we referred to the most recent revision [2]. A haplotype network for N. davidi only was inferred using the Minimum Spanning Network method in PopArt v1.7 [33]. All the sequences generated in this study were submitted to NCBI GenBank [hosts: PV394804–PV394813 (COI), PV697566–PV697569 (12S), PV696985–PV696986 (16S); microsporidians: PV383432–PV383436 (SSU rRNA)].
Statistical analyses
Sex, season, size, and origin of the shrimp individuals might influence parasite infections in N. davidi. However, direct comparisons between these variables were not feasible using a global Generalized Linear Model, as samples were collected in different months with unbalanced sampling effort. Thus, we first compared the sex ratios of sexed N. davidi individuals from different origins (Fontcaude Thermal Park, Mosson River, and pet trade) and separately examined differences across sampling months (March, September, October, and November) using a Chi-square (χ2) test of independence. Then, we compared microsporidian infections between feral and pet traded individuals using Fisher’s exact test. Lastly, we assessed whether the size of sexed individuals influenced microsporidian infections using a logistic regression with infected/uninfected as the dependent variable, and carapax size and sex (female/male) as independent explanatory variables. The results were reported as odds ratios (OR). Statistical and descriptive analyses were performed using the open-source software R v4.4.1 (R Core Team) through the RStudio graphical user interface (version 2024.12.1, Posit Software PBC).
Results
Sample collection
A large number of individuals of N. davidi were observed in the Fontcaude Thermal Park in all sampling months (March, September, and November). Likewise, we confirmed widespread presence of N. davidi along the investigated stretch of the Mosson River in September and November 2024, albeit in smaller numbers than observed in the park. The water parameters measured along the thermal gradient of the Fontcaude Thermal Park at the time of collection were 15.20–23.30 °C, 6.86–7.50 pH, and 3.88–7.53 mg/L DO in March to 21.50–24.00 °C, 7.20–7.66 pH, and 3.37–7.05 mg/L DO in September, and those in the nearby Mosson from 20.30 °C, 7.43 pH, 6.32 mg/L DO in September to 13.80 °C, 7.92 pH, and 9.70 mg/L DO in November.
In total, we collected and examined 320 N. davidi individuals (201 from the thermal park, 67 from the Mosson, and 52 from the pet trade). Field-collected N. davidi comprised 121 females, of which 34 were ovigerous, 125 males, and 22 immature individuals, while pet-traded ones included 24 females, none of which were ovigerous, 22 males, and six immature individuals. Two ovigerous females were detected in the Mosson between September and November 2024. Likewise, immature individuals (carapax size ranging from 1.55 to 2.97 mm) were detected in the Mosson River in both months. No difference in sex ratios was detected among N. davidi individuals from the thermal park, the Mosson, the pet trade, or across sampling months (χ2, all p > 0.28).
Additionally, we collected 126 co-occurring crustaceans from the thermal park and the Mosson which were later molecularly identified as Asellus aquaticus (Linnaeus, 1758) (n = 18, 99.84% similarity and 100% coverage to AY531783), Atyaephyra desmarestii (Millet, 1831) (n = 44, 98.91% similarity and 100% coverage to JX853920), Gammarus pulex (Linnaeus, 1758) (n = 53, 99.15% similarity and 93% coverage to PQ817982), and P. clarkii (n = 11, 100% similarity and coverage to JX441375). These were less widespread than N. davidi, as indicated by visual observations and the number of individuals collected.
Phylogenetic relationships of N. davidi
All feral and pet-traded N. davidi clustered in the clade B1 (Fig. 1). Feral N. davidi belonged to four COI haplotypes, here named “Ndh3”, “Ndh6”, “Ndh7”, and “Ndh09” following [52], and those bought from the pet trade to “Ndh2”, “Ndh3”, “Ndh6”, and “Ndh8” haplotypes (Fig. 2). The most common haplotype was “Ndh6”, which was detected in all samples except those bought from a local physical store. Likewise, “Ndh3” was shared between individuals from the Mosson and those bought from local private sellers. Haplotypes “Ndh9” and “Ndh7” were found exclusively in feral N. davidi individuals, the first in both the thermal park and the Mosson, and the latter only in the Mosson. Conversely, haplotypes “Ndh2” and “Ndh8” were exclusively found in pet-traded individuals; the first was shared between physical and online stores, and the latter was exclusive to a local physical store (Fig. 2).
 |
Figure 1 Maximum likelihood phylogenetic tree of Neocaridina davidi identified in this study. Grey dots represent bootstrap support values above 90%. Sequences obtained in this study are indicated in bold. The substitution model used was HKY + F + I, and N. palmata represents the outgroup. The names and circumscriptions of N. davidi clades followed [52]. Abbreviations: Brazil (BR), Canada (CA), China (CN), Germany (DE), France (FR), Hungary (HU), Israel (IL), Japan (JP), Slovakia (SK), Poland (PL), Taiwan (TW), and the United States of America (USA). |
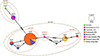 |
Figure 2 Minimum Spanning haplotype network based on COI sequences, including feral and pet traded Neocaridina davidi individuals obtained in this study (in bold) and revisited reference material belonging to three haplotypes (Ndh1, Ndh4, and Ndh5) from Germany, Hungary, and Slovakia, obtained in a previous study (see [52]). Dashed ellipses indicate haplotype circumscriptions based on the newly provided alternative markers 12S and 16S. |
The “Ndh2” haplotype was 100% identical to feral and pet-traded individuals from Brazil (PP718677), Canada (MG319788), Germany, Hungary, Poland, Slovakia (OR610863), the French overseas territory of La Réunion (PQ069719), and wild individuals from Taiwan (MG734262). “Ndh3” showed 100% similarity to feral and pet-traded N. davidi from Germany (OR610861) and wild Taiwanese specimens (MG734280), while “Ndh6” shared 100% similarity with those previously found in the Fontcaude thermal park (PP593877). The haplotypes “Ndh7”, “Ndh8”, and “Ndh9” are reported for the first time in this study (Figs. 1 and 2).
The newly developed primers successfully produced forward sequences of ~600 bp (12S) and ~1,000 bp (16S). These regions were less variable than the COI and resulted in fewer haplotypes. The obtained sequences clustered into four 12S haplotypes and two 16S haplotypes, with divergences of 0.18–0.71% and 0.01%, respectively, further confirming that all analyzed individuals belonged to a single species with a common origin (Fig. 2).
Microsporidian identification and their phylogenetic relationships
In total, 26 out of 320 (8.12%) N. davidi individuals were infected with microsporidians. With only one immature individual infected, the prevalence of microsporidians was higher in adults. Among mature individuals, microsporidian prevalence did not differ between sexes and was not influenced by host size (all ORs 95% CI crossing 1, Table 1). With 24 out of 268 (8.96%) compared to 2 out of 52 (3.85%) infected shrimp, the overall prevalence of microsporidians in feral N. davidi was more than double that of pet-traded individuals. However, this difference was not statistically significant (Fisher’s exact test, p = 0.277). While pet-traded N. davidi (only those from a local physical store) were exclusively infected with E. hepatopenaei (99.70% similarity and 100.00% coverage to KX981865), feral individuals hosted three microsporidian parasites (Fig. 3, Table 2). The most common parasite in feral N. davidi, with a prevalence of 5.60% (15/268), was a newly isolated microsporidian, here named Microsporidium sp. MO02, while the two remaining microsporidians, Microsporidium sp. I and Nosema sp. clade F sensu [2] reached a prevalence of 2.24% (6/268) and 1.12% (3/268), respectively. Microsporidium sp. MO02 and Microsporidium sp. I were shared with other host species, respectively attaining 4.04% (18/446) and 2.24% (10/446) prevalence among all investigated crustaceans. Microsporidium sp. MO02 was only found in adult N. davidi in March 2024 and shared with co-occurring A. aquaticus. This parasite clustered among Gurleyidae in the Amblyosporida order sensu [5], with the most similar isolate being Hazardia milleri (Hazard and Fukuda, 1974) (89.70% similarity and 99.00% coverage to AY090067), isolated from the dipteran Culex quinquefasciatus Say, 1823. Microsporidium sp. I was found in adult N. davidi in September and November 2024 and shared with co-occurring A. aquaticus, G. pulex, and P. clarkii. In contrast, Nosema sp. clade F was only found in immature and adult N. davidi individuals in November. Microsporidium sp. I shared 100.00% similarity and coverage to KR871371, whereas Nosema sp. clade F had 99.63% similarity and 94.00% coverage to MK241529; both reference sequences were isolated from the amphipod G. pulex. Microsporidium sp. MO01, another undescribed isolate, was detected exclusively in co-occurring G. pulex. This parasite clustered with the most similar isolate, Heterovesicula cowani (Lange et al., 1995) (86.48% similarity and 100.00% coverage to EU275200), isolated from the orthopteran Anabrus simplex Haldeman, 1852 among the Heterovesiculidae in the Nosematida order. We found no microsporidian infection in A. desmarestii (Table 2).
 |
Figure 3 Maximum likelihood phylogenetic tree of microsporidians identified in this study. Grey dots represent bootstrap support values above 90%. Sequences obtained in this study are indicated in bold. The substitution model used was TIM3 + F + I + R4, and Metchnikovella dogieli represents the outgroup. The names and circumscriptions of microsporidian orders followed [5]. |
Logistic regression output table from Generalized Linear Model (GLM) showing the influence of carapax size (mm) and host sex (female/male) on microsporidian infection status (infected/uninfected). The number of observations was 292, and Nagelkerke’s R2 was 0.014.
Prevalence of microsporidian in host species collected between March and November 2024.
Discussion
Current status and expansion of N. davidi in European temperate waters
The presence of feral N. davidi populations in continental Europe dates back at least to 2003, with the establishment of a population in northern Poland [22]. In the following years, additional established feral populations have been found in Germany, Hungary, and Slovakia, with individuals reported from both thermal and temperate waters, indicating ongoing expansion into colder waters [30, 52, 71]. More recently, a self-sustaining feral N. davidi population was reported from metropolitan France, at the location investigated in the present study, the Fontcaude Thermal Park [24]. There, the natural warm water flows from the park into the nearby Mosson River [24], where the environmentally adaptable N. davidi appears to have found a suitable habitat. Accordingly, juveniles and ovigerous females were collected in the Mosson during the September and November sampling campaigns, providing further evidence of the species’ expansion into European waters subject to natural thermal regimes.
The water temperature in the Mosson appears to vary between 5.7 °C in winter and 23.5 °C in summer (2008–2015, Naiades, https://naiades.eaufrance.fr), a range that falls within the 6–30 °C reported in N. davidi’s native habitats [30]. Previous observations of ovigerous females at temperatures below 10 °C by [52] suggest that reproduction in the Mosson might even occur during the winter. Habitat suitability predictions from species distribution models indicated a score of 0.6 (low suitability) for the Montpellier area [52], suggesting that the establishment within the local riverine systems of N. davidi, while unlikely, may still be possible. However, these predictions are based on clade B2, which represents northern populations of N. davidi living in colder waters in North China and Japan, and uses air temperature data as a proxy for water temperatures. This clade is phylogenetically and geographically distinct from clade B1, which we found in the present study. Nonetheless, individuals from clade B1, which make up the southern population of N. davidi inhabiting the warmer waters of Taiwan and south China, have also been detected in the colder waters of northern China and Japan [47, 52]. This suggests that both the southern (clade B1) and northern (clade B2) N. davidi populations may have comparable thermal tolerances.
Similarly to other non-native species [36], the presence of a temperature gradient from warm (Fontcaude thermal park) to colder water (Mosson River) might facilitate the adaptation of N. davidi individuals to colder temperatures, a process that is further favored by rising water temperatures, increase and duration of heat waves, and a decrease in cold spells during winter [22, 66, 67, 73]. Alternatively, a large number of N. davidi individuals might drift continuously into the Mosson, facilitating the survival of the population even in the absence of reproduction during the coldest months and/or under predatory pressure. Accordingly, most N. davidi individuals from the thermal park and the Mosson shared identical haplotypes, suggesting genetic flow between the two water bodies.
The role of the pet trade
The presence of haplotypes shared between feral and pet-traded N. davidi clearly indicates a link between the availability of this species in the pet trade and aquarium-dumping practices. Accordingly, the “Ndh3” and “Ndh6” haplotypes were detected in feral and pet-traded individuals bought from local private sellers, suggesting releases of shrimps kept by local aquarists in the thermal park and the Mosson. Furthermore, both “Ndh3” and “Ndh6” haplotypes were also detected in pet-traded individuals bought from one of the largest online stores in Europe (see also [52]). Thus, it is possible that, to a certain degree, the local aquarists sourced their shrimps from online stores. Online stores have increasingly played an influential role in boosting the pet trade by often providing cheap and readily available animals [31]. One of the cheapest and most readily available shrimp species on the online market is N. davidi, which, in its red cherry coloration, can be sold for less than one Euro per individual, significantly lower than the 3–6 Euros usually proposed by physical stores. Since the release likelihood is highest for cheap, prolific, and widely available species [12, 37], the finding of feral N. davidi in the Fontcaude thermal park and the Mosson River is not surprising.
Commercially successful species, including N. davidi, are often mass-produced in aquaculture facilities. High densities in these facilities, coupled with inbreeding, immunosuppression, and suboptimal environmental and nutritional conditions, create an ideal environment for pathogens, such as microsporidian parasites, to proliferate [6, 40, 52, 59]. Several parasites may be lost along the supply chain, for instance, after preventive treatments, handling, and transportation. However, some may still find their way into private aquariums due to insufficient biosecurity measures and the high volume of traded animals [39, 48, 49, 52]. Therefore, with the release of non-native ornamental pets, parasites might be introduced alongside their host into novel environments.
Microsporidian spillover
The current investigation revealed infections with the microsporidian parasite E. hepatopenaei in pet-traded N. davidi individuals bought from a local physical store. This parasite has previously been reported from pet-traded and feral N. davidi in continental Europe and feral individuals in the French overseas territory of La Réunion [51, 52, 56]. Ecytonucleospora hepatopenaei, globally recognized for causing severe economic loss in shrimp aquaculture, has a simple life cycle, is environmentally tolerant, highly contagious, and can infect a wide range of invertebrate hosts, including dragonflies and the highly invasive red swamp crayfish P. clarkii [10, 25, 35, 68]. This parasite is present in pet-traded N. davidi, suggesting that biosecurity measures implemented along the supply chain are still insufficient. Thus, infected N. davidi might still end up in novel environments. However, contrary to our expectations, this parasite was not detected in the feral N. davidi population inhabiting the thermal park or the Mosson.
The absence of any microsporidian parasite clearly originating from the pet trade in the investigated feral N. davidi may indicate a limited subset of introduced animals stemming from uninfected pet-traded individuals. This situation likely hindered spillover events to co-occurring crustaceans. However, it is not excluded that following ongoing aquarium dumping practices, pet-traded pathogens could end up in the investigated ecosystem in the foreseeable future.
Microsporidian spillback
Microsporidium sp. I and Nosema sp. clade F were most likely the result of parasite acquisition from native species. Accordingly, Microsporidium sp. I, also referred to as Microsporidium sp. IV-B in the literature [54] is a host generalist parasite of amphipods that has been detected across Europe in Gammarus balcanicus Schäferna, 1923, G. pulex, G. roeselii Gervais, 1835, and Niphargus schellenbergi Karaman, 1932 [19, 53–55, 70]. Likewise, Nosema sp. clade F is a parasite found in G. pulex and G. balcanicus populations from Central and Meridional Europe, including G. pulex individuals collected from the same region of France where our study was conducted [2]. Moreover, several members of the genus Nosema are known to infect decapods, lepidopterans, neuropterans, and hemipterans [2, 64]. The presence of Nosema sp. clade F in feral N. davidi, and in the case of Microsporidium sp. I also in A. aquaticus and P. clarkii, have not previously been reported. The novel Microsporidium sp. MO02 isolate also seems to be a generalist host parasite. It was shared with A. aquaticus and only detected when A. aquaticus was present in the spring water body, suggesting local origins. Such microsporidians may be able to infect a broader range of species than initially expected, thereby providing a high potential for spillback events. Thus, this reinforces the notion that generalist parasites with a presumably simple life cycle are those that persist better in anthropized environments [53].
The presence of locally acquired host-generalist parasites in feral N. davidi supports our second hypothesis that parasite spillback is favored over spillover. According to the definition of parasite spillback of Kelly et al. [29], when a non-native species functions as an alternative host for native or previously established non-native parasites, it might amplify infection levels in native host populations. For this to happen, the non-native species must acquire native or previously established parasites, be a suitable host in which parasite populations can amplify, attain sufficiently high densities relative to native species so that it can act as a reservoir of infection, and ultimately spillback parasites to the native species [29, 63]. Neocaridina davidi seems to be a suitable alternative host for native or previously established parasites, possibly due to the large number of individuals thriving at the site. Host density is considered a primary driver for transmission rate [1], and several host-generalist microsporidians can switch hosts [60]. Hence, it seems profitable for native or previously established microsporidians to switch from native hosts to the abundant N. davidi, as this will ensure their persistence and enhance their transmission within the ecosystem. Nosema sp. clade F infected specimens of N. davidi, even when the examined native host was not infected. Although we have likely not detected these parasites in native species due to the lower sampling size compared to N. davidi, this finding suggests that N. davidi could act as a reservoir of infection for native microsporidians. Moreover, seasonal dynamics might have influenced the partitioning of parasitic infections among the hosts, as microsporidian prevalence and diversity varied between seasons, supporting our third hypothesis. Importantly, Microsporidium sp. MO01 and MO02 were only detected in March, while Microsporidium sp. I and Nosema sp. clade F were only detected in the fall samples. Therefore, we cannot confirm that parasite spillback has occurred locally with the available data; however, all the prerequisites are present.
Conclusion
All in all, this study provides evidence that non-native, pet-traded species may be suitable alternative hosts for native or previously established host-generalist microsporidian parasites and possess all the prerequisites necessary for parasite spillback events to occur. If spillback events occur, they may impact native host species at both the individual and population levels. Furthermore, our data support the evidence of ongoing range expansion of feral N. davidi into European temperate waters, which is likely bolstered by human-mediated introductions. Therefore, with the ongoing release of pet-traded species into novel environments, particularly in those influenced by warm water, we urge the implementation of preventive measures, including stricter biosecurity measures for pet-traded organisms at all levels of the supply chain, seasonal monitoring of population dynamics in invaded environments, eradication of feral populations, enhanced recognition of responsible pet ownership, and education campaigns tailored to the general public perception of invasive species.
Acknowledgments
This study was performed within the Collaborative Research Center (CRC) RESIST (A09) and funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – CRC 1439/1 – project number: 426547801. We are grateful for the support from the Open Access Publication Fund of the University of Duisburg-Essen. We are very thankful to Ludovic Rossell for his assistance in the field. Lastly, we are very grateful to the anonymous reviewers for their valuable and thorough suggestions that helped to substantially improve the quality of our manuscript.
Conflicts of interest
The authors have no financial or non-financial interests to disclose.
References
- Arneberg P, Skorping A, Grenfell B, Read AF. 1998. Host densities as determinants of abundance in parasite communities. Proceedings of the Royal Society of London. Series B: Biological Sciences, 265, 1283–1289. [Google Scholar]
- Bacela-Spychalska K, Wattier R, Teixeira M, Cordaux R, Quiles A, Grabowski M, Wroblewski P, Ovcharenko M, Grabner D, Weber D, Weigand AM, Rigaud T. 2023. Widespread infection, diversification and old host associations of Nosema Microsporidia in European freshwater gammarids (Amphipoda). PLOS Pathogens, 19, e1011560. [Google Scholar]
- Bauer J, Jung-Schroers V, Teitge F, Adamek M, Steinhagen D. 2021. Association of the alga Cladogonium sp. with a multifactorial disease outbreak in dwarf shrimp (Neocaridina davidi). Diseases of Aquatic Organisms, 146, 107–115. [Google Scholar]
- Bláha M, Weiperth A, Patoka J, Szajbert B, Balogh ER, Staszny Á, Ferincz Á, Lente V, Maciaszek R, Kouba A. 2022. The pet trade as a source of non-native decapods: the case of crayfish and shrimps in a thermal waterbody in Hungary. Environmental Monitoring and Assessment, 194, 795. [Google Scholar]
- Bojko J, Reinke AW, Stentiford GD, Williams B, Rogers MSJ, Bass D. 2022. Microsporidia: a new taxonomic, evolutionary, and ecological synthesis. Trends in Parasitology, 38, 642–659. [CrossRef] [PubMed] [Google Scholar]
- Bojko J, Stentiford GD. 2022. Microsporidian pathogens of aquatic animals, in Microsporidia: Current Advances in Biology, Weiss LM, Reinke AW, Editors. Springer International Publishing: Cham. p. 247–283. [Google Scholar]
- Britton JR, Lynch AJ, Bardal H, Bradbeer SJ, Coetzee JA, Coughlan NE, Dalu T, Tricarico E, Gallardo B, Lintermans M, Lucy F, Liu C, Olden JD, Raghavan R, Pritchard EG. 2023. Preventing and controlling nonnative species invasions to bend the curve of global freshwater biodiversity loss. Environmental Reviews, 31, 310–326. [Google Scholar]
- Chan FT, Beatty SJ, Gilles AS, Hill JE, Kozic S, Luo D, Morgan DL, Pavia RTB, Therriault TW, Verreycken H, Vilizzi L, Wei H, Yeo DCJ, Zeng Y, Zięba G, Copp GH. 2019. Leaving the fish bowl: the ornamental trade as a global vector for freshwater fish invasions. Aquatic Ecosystem Health & Management, 22, 417–439. [Google Scholar]
- Darbyshire AK, Oliver KH, Dupont WD, Plummer WD, Jones CP, Boyd KL. 2019. Anesthesia and euthanasia of brine shrimp (Artemia franciscana). Journal of the American Association for Laboratory Animal Science, 58, 58–64. [Google Scholar]
- Dewangan NK, Pang J, Zhao C, Cao C, Yin B, Weng S, He J. 2023. Host and transmission route of Enterocytozoon hepatopenaei (EHP) between dragonfly and shrimp. Aquaculture, 574, 739642. [Google Scholar]
- Díaz-Morales DM, Sures B, Jolma ER, Thieltges DW. 2025. Invasion biology in the context of aquatic host–parasite interactions, in Aquatic Parasitology: Ecological and Environmental Concepts and Implications of Marine and Freshwater Parasites,Smit NJ, Sures B, Editors. Cham: Springer Nature Switzerland. p. 471–491. [Google Scholar]
- Dickey JWE, Liu C, Briski E, Wolter C, Moesch S, Jeschke JM. 2023. Identifying potential emerging invasive non-native species from the freshwater pet trade. People and Nature, 5, 1948–1961. [Google Scholar]
- Farias L, Beszteri B, Burfeid Castellanos AM, Doliwa A, Enss J, Feld CK, Grabner D, Lampert KP, Mayombo NAS, Prati S, Schürings C, Smollich E, Schäfer RB, Sures B, Le TTY. 2024. Influence of salinity on the thermal tolerance of aquatic organisms. Science of the Total Environment, 953, 176120. [Google Scholar]
- Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. 1994. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology, 3, 294–299. [PubMed] [Google Scholar]
- Gallardo B, Clavero M, Sánchez MI, Vilà M. 2016. Global ecological impacts of invasive species in aquatic ecosystems. Global Change Biology, 22, 151–163. [Google Scholar]
- Gherardi F. 2007. Biological invaders in inland waters: profiles, distribution, and threats. Dordrecht: Springer Netherlands. [Google Scholar]
- Gippet JMW, Bertelsmeier C. 2021. Invasiveness is linked to greater commercial success in the global pet trade. Proceedings of the National Academy of Sciences, 118, e2016337118. [Google Scholar]
- Grabner D, Doliwa A, Sworobowicz L, Wysocka A, Weigand A, Grabowski M, Mamos T, Sures B. 2022. Microsporidian diversity in the aquatic isopod Asellus aquaticus. Parasitology, 149, 1729–1736. [Google Scholar]
- Grabner DS, Weigand AM, Leese F, Winking C, Hering D, Tollrian R, Sures B. 2015. Invaders, natives and their enemies: distribution patterns of amphipods and their microsporidian parasites in the Ruhr Metropolis, Germany. Parasites & Vectors, 8, 419. [CrossRef] [PubMed] [Google Scholar]
- Hassan M, Beatty SJ, Morgan DL, Doupé RG, Lymbery AJ. 2008. An introduced parasite, Lernaea cyprinacea L., found on native freshwater fishes in the south west of Western Australia. Journal of the Royal Society of Western Australia, 91, 149–153. [Google Scholar]
- Havel JE, Kovalenko KE, Thomaz SM, Amalfitano S, Kats LB. 2015. Aquatic invasive species: challenges for the future. Hydrobiologia, 750, 147–170. [CrossRef] [Google Scholar]
- Jabłońska A, Mamos T, Gruszka P, Szlauer-Łukaszewska A, Grabowski M. 2018. First record and DNA barcodes of the aquarium shrimp, Neocaridina davidi, in Central Europe from thermally polluted River Oder canal, Poland. Knowledge & Management of Aquatic Ecosystems, 419, 14. [Google Scholar]
- Jaklič M, Vrezec A. 2011. The first tropical alien crayfish species in european waters: the redclaw Cherax Quadricarinatus (Von Martens, 1868) (Decapoda, Parastacidae). Crustaceana, 84, 651–665. [Google Scholar]
- Jamonneau T, Reyes Camargo AC, Ben Halima I, Prati S, Denys G. 2025. Evidence of an established population of Poecilia reticulata and Neocaridina davidi in metropolitan France. Cybium, in press, https://doi.org/10.26028/cybium/2024-034. [Google Scholar]
- Jang G-I, Kim S-M, Oh Y-K, Lee S-J, Hong S-Y, Lee H-E, Kwon M-G, Kim B-S. 2022. First report of Enterocytozoon hepatopenaei infection in giant freshwater prawn (Macrobrachium rosenbergii de Man) cultured in the Republic of Korea. Animals, 12, 3149. [Google Scholar]
- Kakui K, Komai T. 2022. First record of Scutariella japonica (Platyhelminthes: Rhabdocoela) from Hokkaido, Japan, and notes on its host shrimp Neocaridina sp. aff. davidi (Decapoda: Caridea: Atyidae). Aquatic Animals, AA2022-1. [Google Scholar]
- Karthikeyan K, Sudhakaran R. 2020. Exploring the potentiality of Artemia salina to act as a reservoir for microsporidian Enterocytozoon hepatopenaei of penaeid shrimp. Biocatalysis and Agricultural Biotechnology, 25, 101607. [Google Scholar]
- Katoh K, Rozewicki J, Yamada KD. 2019. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20, 1160–1166. [CrossRef] [PubMed] [Google Scholar]
- Kelly DW, Paterson RA, Townsend CR, Poulin R, Tompkins DM. 2009. Parasite spillback: A neglected concept in invasion ecology? Ecology, 90, 2047–2056. [CrossRef] [PubMed] [Google Scholar]
- Klotz W, Miesen F, Hüllen S, Herder F. 2013. Two Asian freshwater shrimp species found in a thermally polluted stream system in North Rhine-Westphalia, Germany. Aquatic Invasions, 8, 333–339. [Google Scholar]
- Koo KS, Park HR, Choi JH, Sung HC. 2020. Present status of non-native amphibians and reptiles traded in Korean online pet shop. Korean Journal of Environment and Ecology, 34, 106–114. [Google Scholar]
- Krishnan AN, Kannappan S, Aneesh PT, Praveena PE, Jithendran KP. 2021. Polychaete worm – A passive carrier for Enterocytozoon hepatopenaei in shrimp. Aquaculture, 545, 737187. [Google Scholar]
- Leigh JW, Bryant D. 2015. Popart: full-feature software for haplotype network construction. Methods in Ecology and Evolution, 6, 1110–1116. [Google Scholar]
- Liao C-C, Shin J, Chen L-R, Huang LLH, Lin W-C. 2018. First molecular identification of Vorticella sp. from freshwater shrimps in Tainan, Taiwan. International Journal for Parasitology: Parasites and Wildlife, 7, 415–422. [Google Scholar]
- Ling B, Wu Y, Yu Q, Wang C, Hu M, Meng X, Long M, Pan G, Xiang Z, Zhou Z, Chen J. 2024. Ecytonucleospora hepatopenaei proliferate in Procambarus clarkii: A warning for crayfish and shrimp aquaculture. Aquaculture, 581, 740457. [Google Scholar]
- Lipták B, Prati S, Oficialdegui F, Apfelová M, Pekárová S, Kautman J, Janský V, Kouba A. 2024. First populations of invasive red swamp crayfish flourish in Slovakia. BioInvasions Records, 13, 825–841. [Google Scholar]
- Lockwood JL, Welbourne DJ, Romagosa CM, Cassey P, Mandrak NE, Strecker A, Leung B, Stringham OC, Udell B, Episcopio-Sturgeon DJ, Tlusty MF, Sinclair J, Springborn MR, Pienaar EF, Rhyne AL, Keller R. 2019. When pets become pests: the role of the exotic pet trade in producing invasive vertebrate animals. Frontiers in Ecology and the Environment, 17, 323–330. [Google Scholar]
- Maceda-Veiga A, Escribano-Alacid J, De Sostoa A, García-Berthou E. 2013. The aquarium trade as a potential source of fish introductions in southwestern Europe. Biological Invasions, 15, 2707–2716. [Google Scholar]
- Maciaszek R, Świderek W, Kaliszewicz A, Karaban K, Szpakowski B. 2021. First report of Scutariella japonica (Matjašič, 1990), a temnocephalid epibiont from South-East Asia, found on introduced ornamental freshwater shrimp in European waters. Knowledge & Management of Aquatic Ecosystems, 422, 19. [Google Scholar]
- Maciaszek R, Świderek W, Prati S, Huang C-Y, Karaban K, Kaliszewicz A, Jabłońska A. 2023. Epibiont cohabitation in freshwater shrimp Neocaridina davidi with the description of two species new to science, Cladogonium kumaki sp. nov. and Monodiscus kumaki sp. nov., and redescription of Scutariella japonica and Holtodrilus truncatus. Animals, 13, 1616. [Google Scholar]
- McClymont HE, Dunn AM, Terry RS, Rollinson D, Littlewood DTJ, Smith JE. 2005. Molecular data suggest that microsporidian parasites in freshwater snails are diverse. International Journal for Parasitology, 35, 1071–1078. [CrossRef] [PubMed] [Google Scholar]
- Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Haeseler A, von, Lanfear R. 2020. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution, 37, 1530–1534. [CrossRef] [PubMed] [Google Scholar]
- Mrugała A, Kozubíková-Balcarová E, Chucholl C, Cabanillas Resino S, Viljamaa-Dirks S, Vukić J, Petrusek A. 2015. Trade of ornamental crayfish in Europe as a possible introduction pathway for important crustacean diseases: crayfish plague and white spot syndrome. Biological Invasions, 17, 1313–1326. [Google Scholar]
- Munkongwongsiri N, Thepmanee O, Lertsiri K, Vanichviriyakit R, Itsathitphaisarn O, Sritunyalucksana K. 2022. False mussels (Mytilopsis leucophaeata) can be mechanical carriers of the shrimp microsporidian Enterocytozoon hepatopenaei (EHP). Journal of Invertebrate Pathology, 187, 107690. [Google Scholar]
- Niwa N, Ohtaka A. 2006. Accidental introduction of symbionts with imported freshwater shrimps, in Assessment and control of Biological Invasion Risks, Koike F, Clout MN, Kawamichi M, De Poorter M, Iwatsuki K, Editors. Cambridge, UK: IUCN. p. 182–186. [Google Scholar]
- Ohtaka A, Gelder SR, Nishino M, Ikeda M, Toyama H, Cui Y-D, He X-B, Wang H-Z, Chen R-B, Wang Z-Y. 2012. Distributions of two ectosymbionts, branchiobdellidans (Annelida: Clitellata) and scutariellids (Platyhelminthes: “Turbellaria”: Temnocephalida), on atyid shrimp (Arthropoda: Crustacea) in southeast China. Journal of Natural History, 46, 1547–1556. [Google Scholar]
- Onuki K, Fuke Y. 2022. Rediscovery of a native freshwater shrimp, Neocaridina denticulata, and expansion of an invasive species in and around Lake Biwa, Japan: genetic and morphological approach. Conservation Genetics, 23, 967–980. [Google Scholar]
- Patoka J, Bláha M, Devetter M, Rylková K, Čadková Z, Kalous L. 2016. Aquarium hitchhikers: attached commensals imported with freshwater shrimps via the pet trade. Biological Invasions, 18, 457–461. [Google Scholar]
- Patoka J, Magalhães ALB, Kouba A, Faulkes Z, Jerikho R, Vitule JRS. 2018. Invasive aquatic pets: failed policies increase risks of harmful invasions. Biodiversity and Conservation, 27, 3037–3046. [Google Scholar]
- Prati S, Enß J, Grabner DS, Huesken A, Feld CK, Doliwa A, Sures B. 2023. Possible seasonal and diurnal modulation of Gammarus pulex (Crustacea, Amphipoda) drift by microsporidian parasites. Scientific Reports, 13, 9474. [Google Scholar]
- Prati S, Faivre L, de Mazancourt V, Sures B, Valade P. 2025. Paradise under threat: the successful invasion of the freshwater shrimp Neocaridina davidi (Bouvier, 1904) and its microsporidian parasites on La Réunion Island. BioInvasions Records, 14, 403–419. [Google Scholar]
- Prati S, Grabner DS, Hupało K, Weiperth A, Maciaszek R, Lipták B, Bojko J, Bérces F, Sures B. 2024. Invisible invaders: range expansion of feral Neocaridina davidi offers new opportunities for generalist intracellular parasites. Biological Invasions, 26, 2499–2523. [Google Scholar]
- Prati S, Grabner DS, Pfeifer SM, Lorenz AW, Sures B. 2022. Generalist parasites persist in degraded environments: a lesson learned from microsporidian diversity in amphipods. Parasitology, 149, 973–982. [Google Scholar]
- Quiles A, Bacela-Spychalska K, Teixeira M, Lambin N, Grabowski M, Rigaud T, Wattier RA. 2019. Microsporidian infections in the species complex Gammarus roeselii (Amphipoda) over its geographical range: evidence for both host–parasite co-diversification and recent host shifts. Parasites & Vectors, 12, 327. [Google Scholar]
- Quiles A, Rigaud T, Wattier RA, Grabowski M, Bacela Spychalska K. 2021. Wide geographic distribution of overlooked parasites: Rare Microsporidia in Gammarus balcanicus, a species complex with a high rate of endemism. International Journal for Parasitology: Parasites and Wildlife, 14, 121–129. [Google Scholar]
- Schneider R, Prati S, Grabner D, Sures B. 2022. First report of microsporidians in the non-native shrimp Neocaridina davidi from a temperate European stream. Diseases of Aquatic Organisms, 150, 125–130. [Google Scholar]
- Schoolmann G, Arndt H. 2017. Leaf-litter preferences of the introduced freshwater shrimps Atyaephyra desmarestii and Neocaridina davidi. Crustaceana, 90, 1715–1730. [Google Scholar]
- Simberloff D, Martin J-L, Genovesi P, Maris V, Wardle D, Aronson J, Courchamp F, Galil B, García-Berthou E, Pascal M, Pyšek P, Sousa R, Tabacchi E, Vilà M. 2013. Impacts of biological invasions: what’s what and the way forward. Trends in Ecology & Evolution, 28, 58–66. [CrossRef] [PubMed] [Google Scholar]
- Smallbone W, van Oosterhout C, Cable J. 2016. The effects of inbreeding on disease susceptibility: Gyrodactylus turnbulli infection of guppies, Poecilia reticulata. Experimental Parasitology, 167, 32–37. [Google Scholar]
- Stentiford GD, Feist SW, Stone DM, Bateman KS, Dunn AM. 2013. Microsporidia: diverse, dynamic, and emergent pathogens in aquatic systems. Trends in Parasitology, 29, 567–578. [CrossRef] [PubMed] [Google Scholar]
- Strauss A, White A, Boots M. 2012. Invading with biological weapons: the importance of disease-mediated invasions. Functional Ecology, 26, 1249–1261. [Google Scholar]
- Svoboda J, Mrugała A, Kozubíková-Balcarová E, Petrusek A. 2017. Hosts and transmission of the crayfish plague pathogen Aphanomyces astaci: a review. Journal of Fish Diseases, 40, 127–140. [Google Scholar]
- Telfer S, Bown K. 2012. The effects of invasion on parasite dynamics and communities. Functional Ecology, 26, 1288–1299. [Google Scholar]
- Tokarev YS, Huang W-F, Solter LF, Malysh JM, Becnel JJ, Vossbrinck CR. 2020. A formal redefinition of the genera Nosema and Vairimorpha (Microsporidia: Nosematidae) and reassignment of species based on molecular phylogenetics. Journal of Invertebrate Pathology, 169, 107279. [Google Scholar]
- Torchin ME, Lafferty KD, Dobson AP, McKenzie VJ, Kuris AM. 2003. Introduced species and their missing parasites. Nature, 421, 628–630. [CrossRef] [PubMed] [Google Scholar]
- Tuckett QM, Lawson KM, Lipscomb TN, Hill JE, Daniel WM, Siders ZA. 2021. Non-native poeciliids in hot water: the role of thermal springs in facilitating invasion of tropical species. Hydrobiologia, 848, 4731–4745. [Google Scholar]
- Van Vliet MTH, Franssen WHP, Yearsley JR, Ludwig F, Haddeland I, Lettenmaier DP, Kabat P. 2013. Global river discharge and water temperature under climate change. Global Environmental Change, 23, 450–464. [Google Scholar]
- Wan Sajiri WMH, Kua BC, Borkhanuddin MH. 2023. Detection of Enterocytozoon hepatopenaei (EHP) (microsporidia) in several species of potential macrofauna-carriers from shrimp (Penaeus vannamei) ponds in Malaysia. Journal of Invertebrate Pathology, 198, 107910. [Google Scholar]
- Weber S, Traunspurger W. 2016. Influence of the ornamental red cherry shrimp Neocaridina davidi (Bouvier, 1904) on freshwater meiofaunal assemblages. Limnologica, 59, 155–161. [Google Scholar]
- Weigand AM, Kremers J, Grabner DS. 2016. Shared microsporidian profiles between an obligate (Niphargus) and facultative subterranean amphipod population (Gammarus) at sympatry provide indications for underground transmission pathways. Limnologica, 58, 7–10. [CrossRef] [Google Scholar]
- Weiperth A, Gábris V, Danyik T, Farkas A, Kuříková P, Kouba A, Patoka J. 2019. Occurrence of non-native red cherry shrimp in European temperate waterbodies: a case study from Hungary. Knowledge & Management of Aquatic Ecosystems, 420, 9. [Google Scholar]
- Wood LE, Guilder J, Brennan ML, Birland NJ, Taleti V, Stinton N, Taylor NGH, Thrush MA. 2022. Biosecurity and the ornamental fish trade: A stakeholder perspective in England. Journal of Fish Biology, 100, 352–365. [Google Scholar]
- Woolway RI, Jennings E, Shatwell T, Golub M, Pierson DC, Maberly SC. 2021. Lake heatwaves under climate change. Nature, 589, 402–407. [Google Scholar]
- Zhu X, Wittner M, Tanowitz HB, Kotler D, Cali A, Weiss LM. 1993. Small Subunit rRNA sequence of Enterocytozoon bieneusi and its potential diagnostic role with use of the Polymerase Chain Reaction. Journal of Infectious Diseases, 168, 1570–1575. [CrossRef] [PubMed] [Google Scholar]
Cite this article as: Prati S, Reyes Camargo ACR, Jamonneau T, Halima IB & Sures B. 2025. Seasonal exchange of microsporidian parasites between native and non-native pet-traded freshwater crustaceans: Is parasite spillover favored over spillback?. Parasite 32, 61. https://doi.org/10.1051/parasite/2025053.
All Tables
Logistic regression output table from Generalized Linear Model (GLM) showing the influence of carapax size (mm) and host sex (female/male) on microsporidian infection status (infected/uninfected). The number of observations was 292, and Nagelkerke’s R2 was 0.014.
Prevalence of microsporidian in host species collected between March and November 2024.
All Figures
 |
Figure 1 Maximum likelihood phylogenetic tree of Neocaridina davidi identified in this study. Grey dots represent bootstrap support values above 90%. Sequences obtained in this study are indicated in bold. The substitution model used was HKY + F + I, and N. palmata represents the outgroup. The names and circumscriptions of N. davidi clades followed [52]. Abbreviations: Brazil (BR), Canada (CA), China (CN), Germany (DE), France (FR), Hungary (HU), Israel (IL), Japan (JP), Slovakia (SK), Poland (PL), Taiwan (TW), and the United States of America (USA). |
| In the text | |
 |
Figure 2 Minimum Spanning haplotype network based on COI sequences, including feral and pet traded Neocaridina davidi individuals obtained in this study (in bold) and revisited reference material belonging to three haplotypes (Ndh1, Ndh4, and Ndh5) from Germany, Hungary, and Slovakia, obtained in a previous study (see [52]). Dashed ellipses indicate haplotype circumscriptions based on the newly provided alternative markers 12S and 16S. |
| In the text | |
 |
Figure 3 Maximum likelihood phylogenetic tree of microsporidians identified in this study. Grey dots represent bootstrap support values above 90%. Sequences obtained in this study are indicated in bold. The substitution model used was TIM3 + F + I + R4, and Metchnikovella dogieli represents the outgroup. The names and circumscriptions of microsporidian orders followed [5]. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


