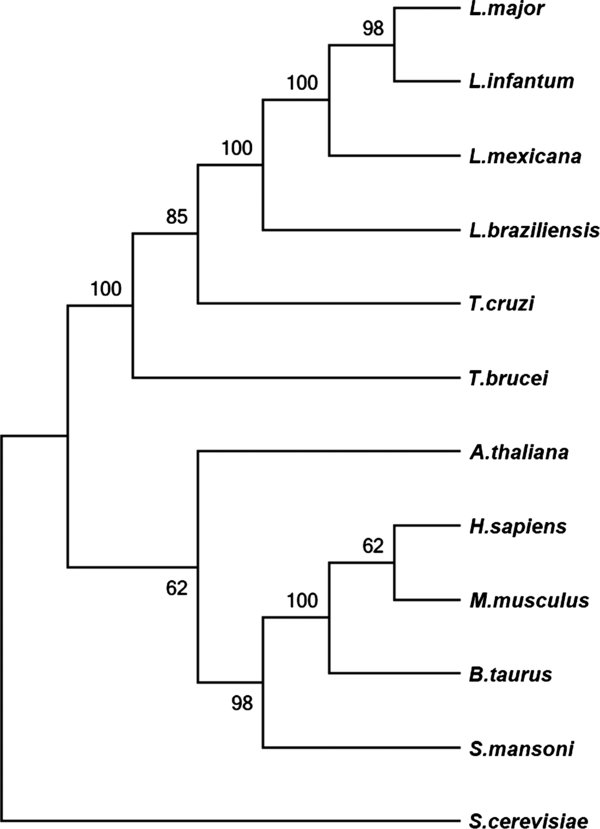
Figure 1.
Download original image
Bioinformatics analysis. The relationship of Leishmania major PP5 (XP_001682421.1) to PP5 homologs of Leishmania infantum (XP_001464832.1), Leishmania mexicana (XP_003874029.1), Leishmania braziliensis (XP_001563942.2), Trypanosoma cruzi (XP_820611.1), Trypanosoma brucei (XP_827850.1), Arabidopsis thaliana (NP_565985.1), Homo sapiens (NP_006238.1), Mus musculus (NP_035285.2), Bos taurus (NP_001179178.1), Schistosoma mansoni (XP_002580099.1) and Saccharomyces cerevisiae (EDN61714.1) was analyzed by multiple alignment and cluster analysis using Clustal X. Alignment was fed into MEGA5.2 software and a Neighbor-Joining tree was computed with 500 bootstrap replicates. Numbers on nodes indicate bootstrap support.
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


