| Issue |
Parasite
Volume 33, 2026
|
|
|---|---|---|
| Article Number | 15 | |
| Number of page(s) | 16 | |
| DOI | https://doi.org/10.1051/parasite/2026016 | |
| Published online | 30 March 2026 | |
Research Article
Microphallus pseudopygmaeus (Digenea) infects phylogenetically distant gastropods, with signs of host-linked genetic divergence
Microphallus pseudopygmaeus (Digenea) infecte des gastéropodes phylogénétiquement éloignés, avec des signes de divergence génétique liée à l’hôte
1
Department of Invertebrate Zoology, Saint Petersburg State University, Saint Petersburg, Russia
2
Laboratory of Parasitic Worms and Protists, Zoological Institute of the Russian Academy of Sciences, Universitetskaya emb. 1, Saint Petersburg 199034, Russia
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
; This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
6
August
2025
Accepted:
16
February
2026
Abstract
Host-switching between distantly related host species offers rare insight into how parasites overcome compatibility barriers and initiate evolutionary divergence. Microphallus pseudopygmaeus is exceptional among digeneans in its ability to infect gastropods from two distantly related subclasses, Vetigastropoda and Caenogastropoda. This study aimed to test the hypothesis about the species status of M. pseudopygmaeus and clarify its host range. We obtained partial sequences of the cox1 gene, 28S rDNA and ITS2 for M. pseudopygmaeus from nine host species, including Margarites spp. (Vetigastropoda). The data on the conservative and variable markers, phylogenetic and barcoding gap analyses, supported the unity of the species and its broad specificity to the first intermediate hosts. The cox1-based haplotype network revealed host-associated genetic divergence, particularly in isolates from Margarites spp. and Cryptonatica affinis. This pattern may result from localized circulation of the parasite in the regions where certain host species, such as Margarites spp., dominate in the absence of periwinkles, creating ecological conditions that could promote reproductive isolation and incipient speciation. This work opens up the prospects of using M. pseudopygmaeus as a model for studying host-switching and speciation in parasites.
Résumé
Le passage d’un hôte à un autre entre des espèces hôtes éloignées offre un aperçu rare de la façon dont les parasites surmontent les barrières de compatibilité et initient une divergence évolutive. Microphallus pseudopygmaeus est exceptionnel parmi les digènes par sa capacité à infecter des gastéropodes appartenant à deux sous-classes éloignées, les Vetigastropoda et les Caenogastropoda. Cette étude visait à tester l’hypothèse concernant le statut d’espèce de M. pseudopygmaeus et à préciser son spectre d’hôtes. Nous avons obtenu des séquences partielles du gène cox1, de l’ADNr 28S et de l’ITS2 pour M. pseudopygmaeus chez neuf espèces d’hôtes, dont Margarites spp. (Vetigastropoda). Les données sur les marqueurs conservés et variables, l’analyse phylogénétique et l’analyse du barcode gap, ont confirmé l’unité de l’espèce et sa large spécificité envers les premiers hôtes intermédiaires. L’analyse du réseau d’haplotypes basé sur le gène cox1 a révélé une divergence génétique liée à l’hôte, notamment chez les isolats provenant de Margarites spp. et de Cryptonatica affinis. Ce schéma pourrait résulter d’une circulation localisée du parasite dans les régions où certaines espèces hôtes, comme Margarites spp., dominent en l’absence de littorines, créant ainsi des conditions écologiques susceptibles de favoriser l’isolement reproductif et une spéciation naissante. Ces travaux ouvrent la voie à l’utilisation de M. pseudopygmaeus comme modèle d’étude du changement d’hôte et de la spéciation chez les parasites.
Key words: Host specificity / Host switching / First intermediate host / Intraspecific divergence / Speciation
Edited by Jean-Lou Justine
© A. Sokolova et al., published by EDP Sciences, 2026
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Introduction
Parasites pose several challenging questions in evolutionary biology, to name just a few – the origins of parasitism, the rationale for life cycle complexity, the nature of virulence dynamics, the force balancing specificity and generalism. The question of how parasites may retain specificity while rapidly responding to changes is intriguing, and it is currently considered that the primary mode of parasite evolution is not co-speciation but rather host-switching [14, 40]. The latter must be particularly frequent in parasites featuring multi-host life cycles, perfectly exemplified by digeneans.
In the life cycles of these parasitic flatworms, parthenogenetic (asexual) generations (sporocysts and/or rediae) develop in the first intermediate hosts, metacercariae in the environment or in the second intermediate hosts, and maritae (sexual adults) in the definitive hosts. Colonization of new hosts has been the main driver of digenean evolution [5, 8, 18, 32]. Switching definitive hosts can be a hallmark of new lineages, e.g. family Brachycladiidae (fishes to cetaceans, [22]); a major divergence event within lineages, e.g., in Schistosomatidae (birds to mammals, [8]); or a repeated event contributing to a wide host range, e.g., in Cyathocotylidae [1]. Switching second intermediate hosts is also one of the pathways for digeneans to diversify and reach definitive hosts with varying feeding preferences (e.g., in Opecoelidae, [59], Hemiuridae [51]). Evident history of switching first intermediate hosts by digeneans [5, 8, 58, 80] goes along with their remarkable specificity at the fine taxonomic level.
One digenean species is usually restricted to one molluscan genus (e.g., most schistosomes [8], Opisthorchis felineus [61], Podocotyle atomon [50]), or even species (e.g., Himasthla elongata [35], Catatropis onobae [37], Neophasis annarichae [47], Orthosplanchnus arcticus [46, 49]). In some cases, the range of suitable first intermediate hosts is wider and includes multiple members of the same family [23] or even superfamily [52]. Such cases of broader specificity can provide insights into how the colonization of new taxa of first intermediate hosts can drive speciation in digeneans. As a model for this type of study, we propose the species Microphallus pseudopygmaeus Galaktionov, 1980 (Digenea: Microphallidae).
Microphallus pseudopygmaeus belongs to a group of closely related species referred to as the “pygmaeus” group [31]. These digeneans have no free-living cercariae, and metacercariae (Fig. 1) develop directly within daughter sporocysts in the first intermediate host. The life cycle therefore includes only the definitive hosts (marine anatids, mainly eiders; or gulls) and the first intermediate hosts (shared by all members of the “pygmaeus” group are periwinkles, Littorina spp.) (Fig. 2). However, M. pseudopygmaeus has expanded its range of first intermediate hosts to as many as 17 gastropod species, some of which are phylogenetically distant. This conclusion was initially based on morphological observations [25, 28, 29] and later supported by sequencing of the ITS1, ITS2 and 28S rDNA regions [31]. To fully appreciate the host range of M. pseudopygmaeus, we have now expanded the dataset to include the widest available diversity of hosts, and the variable mtDNA marker. The results inspired us to discuss biogeography, host switching and speciation, and envision further studies.
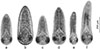 |
Figure 1 Metacercariae of microphallids of the “pygmaeus” group, microphotographs. (a) M. pseudopygmaeus, (b) M. pygmaeus, (c) M. triangulatus, (d) M. kurilensis, (e) M. piriformes, (f) M. calidris. |
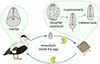 |
Figure 2 Life cycle of microphallids of the “pygmaeus” group. Definitive hosts: marine birds; first intermediate hosts: marine gastropods (typically, periwinkles); trophic transmission: the Pac-Man icon. |
Material and methods
Sampling
Sampling took place on the shores of the northern European seas (Norwegian, Barents, White, Pechora) and the Sea of Okhotsk (Table 1, Fig. 3) in 2003–2025. In addition, we used 10 isolates of microphallids collected by Georgii Kremnev and Darya Krupenko in 2023–2024. All the samples were from the gastropod first intermediate hosts (Table 2). Three periwinkle species belonging to the “saxatilis” complex (Littorina saxatilis, Littorina arcana and Littorina compressa) co-occur at our Barents Sea sampling location [38]. They may be difficult to distinguish when the snail is castrated due to trematode infection. In these cases, we listed them as belonging to the “saxatilis” species complex. During low tide, we collected littoral snails, while the mollusks Cryptonatica affinis (Gmelin, 1791) and Margarites spp. were gathered from the sublittoral zone by dredging and diving. We kept mollusks in containers filled with sea water, and then dissected them to detect infection. Microphallid sporocysts and metacercariae were identified and preserved in 96% ethanol. In addition to M. pseudopygmaeus, we sampled other members of the “pygmaeus” group (Microphallus pygmaeus (Levinsen, 1881) Baer, 1944 [55], Microphallus piriformes Galaktionov, 1983 [26], M. pseudopygmaeus, Microphallus triangulatus Galaktionov, 1984 [27], Microphallus calidris Belopolskaja and Ryzhikov [4], Microphallus kurilensis Galaktionov, Regel and Atrashkevich, 2010 [34]; Fig. 1) and Microphallus similis (Jägerskiöld, 1900) Baer, 1944 [42].
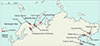 |
Figure 3 Map of all sampling sites (indicated by arrows) and M. pseudopygmaeus sampling sites (red dots). Closely spaced localities (e.g., within the Pechora Sea and the White Sea) are not shown on the map. |
Geographical locations of sampling sites.
Material used in this study and the corresponding GenBank accession numbers.
To identify microphallids of the “pygmaeus” group, we prepared wet mounts as previously described [25, 69, 71]. Slides with metacercariae in a drop of distilled water were heated for ca. 1 min at 70 °C using a heating table. Then, completely relaxed and flattened metacercariae were identified based on morphological characters, and the reference photographs were made using an Olympus CH40 compound microscope (Olympus Optical Co. Ltd., Tokyo, Japan) equipped with an Olympus XC-30 digital camera (Olympus Optical Co. Ltd.) (Fig. 1). We did not use other morphological methods because comprehensive data about the structure of M. pseudopygmaeus metacercariae from different snail hosts are already available [28, 29], and here we focused on identifying intra- and interspecific genetic variability.
The material was preserved in 96% ethanol for further studies. We deposited voucher tissue samples (paragenophores) to the Trematoda voucher collection of the Zoological Institute of the Russian Academy of Science (ZISP) under accession numbers 2025.11.12.001–2025.11.12.114; their correspondence to the GenBank accession numbers of the sequences is provided in Table 2.
Nucleotide sequence data reported in this paper are available in the GenBank database under the accession numbers PQ611031–PQ611083, PQ728074–PQ728079, PQ836125–PQ836130, PQ836132–PQ836137, PV789575–PV789581 and PX560768–PX560778.
DNA extraction, PCR and sequencing
We removed a single sporocyst from ethanol, placed it in a new 1.5-mL microtube, and let any remaining ethanol evaporate by keeping a tube open for 1–3 min. To extract DNA, we added 200 μL of the 5% ion exchange resin Chelex 100 (Bio-Rad, Hercules, CA, USA) and 2 μL of proteinase K (Evrogen, Moscow, Russia) to each tube, then incubated samples at 56 °C for 4 hours while shaking at 850 rpm on a thermomixer (BioSan, Riga, Latvia). To inactivate the proteinase, the samples were then boiled at 90 °C for 8 min. After that, we centrifuged the tubes for 10 min at 16,874 ×g (Eppendorf 5418). The DNA in the supernatant was transferred to a new tube and stored at −20 °C.
To amplify the partial cox1 gene, 5.8S–ITS2 region and D1–D3 domains of 28S rRNA gene, we used the primers listed in Table 3. The primer JB3 was modified based on the draft mitogenome assembly of M. pseudopygmaeus (unpublished data by K.V. Galaktionov) to enhance PCR outcome.
PCR primers used in this study.
Amplification was performed in 20 μL reaction mixtures containing 2 μL DNA template, 0.5 μL forward and reverse primer each (10 pmol/μL), 4 μL ScreenMix-HS reaction mix (Evrogen, Russia), and 13 μL Super-Q water.
PCR conditions for the cox1 fragment were as following: initial denaturation at 95 °C for 5 min, 35 cycles (95 °C – 30 s, 48 °C – 30 s, 72 °C – 1 min), and final elongation at 72 °C for 10 min (T100 BioRad). To amplify the fragment of 5.8S–ITS2, we used the protocol from [47]. To amplify the D1–D3 domains of the 28S rDNA, we used the touchdown PCR protocol with a sequential decrease in the annealing temperature of 55.5 °C (10 cycles) – 55 °C (10 cycles) – 54.5 °C (15 cycles). PCR-products were size-separated by electrophoresis in a 1% agarose gel (SE-1, Helicon), stained with ethidium bromide and photographed in the UV light using ChemiDoc BioRad.
Sanger sequencing (forward and reverse) was performed on an automated capillary sequencer ABI 3500xl (Applied Biosystems, Foster City, CA, USA) at the Centre for Molecular and Cell Technologies, St Petersburg University.
Data processing and analysis
Sequence data were processed in Geneious Prime 23.2.1 [43]. To estimate genetic distances, we used Mega 11 [75]. For new rDNA sequences and those publicly available from GenBank, we calculated mean intra- and interspecific distances with the maximum likelihood estimation method [75]. For all specimens of the “pygmaeus” group microphallids and M. similis, we calculated pairwise p-distances in the cox1 fragment, and used these data to build a histogram of distance frequencies in R [65] and RStudio IDE version 2024.12.0 [62] with a ggplot2 package [81].
Species delimitation hypotheses were tested based on cox1 sequence data using assemble species by automatic partitioning (ASAP) [63] implemented in the iTaxoTools toolkit, with simple distances and default parameters [77]. For the cox1-based phylogenetic reconstruction, we used M. similis as an outgroup and microphallids of the “pygmaeus” group as an ingroup; identical sequences were removed from the alignment. Bayesian inference (BI) was performed in MrBayes, v. 3.2.7a [70] with GTR + inv + G model for 1,000,000 generations (sampling and diagnosing frequencies 1,000, 25% burnout), making sure the runs converged. For the maximum likelihood approach (ML), we used RAxML-NG v. 1.2.2 [44] implemented in raxmlGUI 2.0.13 [20] with the integrated model selection (HKY + FO + I+G4m), “ML + transfer bootstrap expectation + consensus” option, one run, and 100 replicates. The BI tree was visualized, and the poorly supported nodes (posterior probabilities below 0.75) were collapsed into polytomies. For nodes that were also recovered in the ML analysis, both BI and ML support values were printed. ASAP scores were visualized and color-coded next to the resulting tree.
We also created a cox1-haplotype network in PopART 1.7 with the TCS algorithm [13].
For species in the “pygmaeus” group, we also searched for the robust diagnostic nucleotide combinations: combinations of nucleotides at specified sites of the alignment, unique for a particular species and sufficient to differentiate it from all other taxa in a dataset. This search was performed in MolD [21] implemented in iTaxoTools [77].
We prepared the figures in CorelDRAW Graphics Suite, v. 24.0.0.301 [16].
Results
Nuclear markers
We obtained eleven 28S rDNA and thirteen 5.8S–ITS2 rDNA sequences for isolates of M. pseudopygmaeus from different molluscan hosts: L. saxatilis, L. obtusata, Lacuna vincta, O. aculeus, C. affinis, M. helicinus, and M. groenlandicus. These data were analyzed together with the sequences of M. pseudopygmaeus and six other Microphallus species available in GenBank (HM584122–HM584142, HM584175–HM584199, [31]; MG783583–MG783584, MG783588–MG783589, [30]; AY220625, [76]).
The trimmed 28S rDNA alignment was 971 bp long and included 29 sequences. There were no intraspecific variations except for one M. piriformes sequence differing by one substitution. The distances between the species of the “pygmaeus” group (the number of base substitutions per site) were 0–0.034 (Supplementary Table S1). The trimmed 5.8S–ITS2 rDNA alignment was 266 bp long and included 36 sequences. There were also no intraspecific variations except for two sequences of M. pseudopygmaeus from C. affinis which differed by one substitution. The distances between the species of the “pygmaeus” group were 0.06–0.027 (Supplementary Table S2).
Mitochondrial marker cox1
We obtained 104 new cox1 sequences, their length ranging from 795 to 963 bp (M. pseudopygmaeus – 64, M. pygmaeus – 8, M. triangulatus – 2, M. kurilensis – 3, M. piriformes – 11, M. calidris – 1, M. similis – 15). The reverse primer trem.cox1.rrnl flanks a region outside the cox1 gene; this is a non-coding fragment directly adjacent to the cox1 CDS, followed by tRNA. To be on the safe side, for GenBank we annotated the cox1 CDS only and left the remaining fragment unannotated.
Species delimitation
In the cox1-based phylogeny (801 bp alignment including 42 sequences), each microphallid of the “pygmaeus” group was recovered as a well-supported clade (Fig. 4a). This matches well the results of the cox1-based ASAP analysis (801 bp alignment including 104 sequences): the minimum ASAP score (2.0), and thus the highest probability, was assigned to the partition including seven subsets (Fig. 4b). These subsets correspond to the seven assumed species in the alignment, supporting the hypothesis that M. pseudopygmaeus is a single species.
 |
Figure 4 Phylogenetic relationships of the “pygmaeus” group microphallids based on cox1 sequences. (a) Bayesian phylogenetic tree. Poorly supported nodes (posterior probability < 0.75) were collapsed into polytomies. For nodes also recovered in the maximum likelihood (ML) analysis, both BI and ML support values are shown. (b) Five best ASAP scores and partition numbers (in brackets); and color-coded putative species boundaries corresponding to the clades in the tree. |
The boundary value of p-distances was 0.0631, and the barcoding gap is visualized in Figure 5. Most of the pairwise p-distances within M. pseudopygmaeus ranged from 0.00 to 0.014, with five outlier specimens displaying distances above this range, up to 0.056. Intraspecific distances among microphallids of the “pygmaeus” group were above 0.079, and mostly above 0.010. However, there was one exception: M. pygmaeus and M. kurilensis specimens differed by just 0.048–0.051; despite distances below the threshold value, these species are still consistently recognized as distinct in the ASAP analysis (Fig. 4b). Distances between M. similis and the “pygmaeus” group specimens were 0.140–0.156. The complete cox1 distance matrix is provided in Supplementary Table S3; the alignment used for this matrix and ASAP analysis is in Supplementary File.
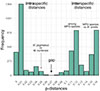 |
Figure 5 Frequency distribution of pairwise genetic distances in the cox1 gene fragment (803 bp) for seven species of Microphallus. MPG, microphallids of the “pygmaeus” group. |
Diagnostic nucleotide combinations (DNCs) were identified in MolD for all members of the “pygmaeus” group microphallids, based on the 801 bp cox1 alignment as used in ASAP analysis, and were as follows: C188 and A191 in M. pygmaeus; C143 and T191 in M. kurilensis; C2 and A29 in M. calidris; C3, T8 and G188 in M. piriformes; C8 and C65 in M. triangulatus; and C188 and A191 in M. pseudopygmaeus. When the analysis was run for the M. pseudopygmaeus isolates subdivided by host (Cryptonatica affinis, Margarites spp. and all other), DNCs were only recovered for isolates from C. affinis: A 35, T38, and C80.
Haplotype network
The alignment contained 74 cox1 sequences: 64 for M. pseudopygmaeus from nine species of molluscan hosts and five each for M. pygmaeus and M. similis to exemplify interspecific distances. The total length of the alignment was 800 bp.
We discovered 28 haplotypes among the putative M. pseudopygmaeus samples. Most of the haplotypes differed by one substitution. There was a minimum of 83 substitutions between M. pseudopygmaeus and the closely related species M. pygmaeus; and 109 between M. pseudopygmaeus and M. similis.
The distribution of M. pseudopygmaeus haplotypes among the species of molluscan hosts is illustrated with colors in Figure 6a. The dominant haplotype A included 27 samples from L. saxatilis, L. obtusata, Lacuna vincta and Onoba aculeus. Two samples of M. pseudopygmaeus from Cryptonatica affinis had the most divergent haplotypes and were only moderately similar to each other (B); they differed from the dominant haplotype A by 30–38 substitutions. Samples of M. pseudopygmaeus from Margarites spp. (M. helicinus and M. groenlandicus) grouped together (haplogroup C); they differed from the dominant haplotype A by 6–10 mutations. Sequences of the samples from L. sitkana formed two haplotypes which differed markedly from each other: one (D) was close to the dominant haplotype (differs by four mutations), while the second one (E) stood alone (differed by 25 substitutions from the dominant haplotype A).
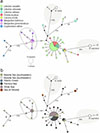 |
Figure 6 Haplotype networks for M. pseudopygmaeus, also including several samples of M. pygmaeus and M. similis, based on the fragment of the cox1 gene (800 bp). Circle sizes represent haplotype frequencies, hatch marks represent substitutions (if more than five, replaced with numbers). The asterisk marks the sample from L. arcana (belongs to “saxatilis” species complex). Haplotypes are colored according to parasite host species (a) and sampling location (b). The sampling locations in the Barents Sea: Dalniye Zelentsy (southwestern), Chosha Bay, Kanin Cape (southeastern); in the Pechora Sea: Dolgiy Island, Vaygach Island, Malyi Zincovyi Island, Gubistyi Island; in the White Sea: Levin Navolok Bay, Cape Kindo, Korovya Varakka Cliff; in the Sea of Okhotsk: Tyuleniy Island, Taygonos Cape, Nagaev Bay; in the Atlantic Ocean: Grindavik. |
The distribution of M. pseudopygmaeus haplotypes among the sampling regions is illustrated with colors in Figure 6b. The genetically distinct haplotype E included the geographically distinct specimens from the North Pacific (Sea of Okhotsk). However, one specimen from this region appeared separately as haplotype D. The dominant haplotype A was shared by specimens originating from all the sampled regions in the North Atlantic. However, one specimen from the North Atlantic (haplotype F, Pechora Sea) appeared relatively closer to haplotype E of North Pacific origin.
Discussion
Microphallus pseudopygmaeus has been described as a species able to infect an extremely wide range of molluscan first intermediate hosts [24, 29]. Later, its host range was partially confirmed with rDNA sequence data [31]. However, considering the occurrence of cryptic and pseudocryptic species among trematodes, the integrity of M. pseudopygmaeus required revision with a variable genetic marker. In this study, we focused on the cox1 gene data and analyzed additional evidence that M. pseudopygmaeus is indeed a single species.
Species delimitation in the microphallids of the “pygmaeus” group
Results of the phylogenetic and ASAP analyses match the previous assumption that “pygmaeus” group comprises six species: M. pygmaeus, M. piriformes, M. pseudopygmaeus, M. triangulatus, M. calidris and M. kurilensis. This implies the unity of M. pseudopygmaeus, with all its specimens from various first intermediate host species. Phylogenetic relationships suggest that it is a sister species to M. triangulatus (Fig. 4), similarly to the previous rDNA-based findings [31]. There is no doubt that M. pseudopygmaeus and M. triangulatus are distinct species, considering genetic (Supplementary Table S3) and morphological (Fig. 1) differences. The distinction between the sister species M. pygmaeus and M. kurilensis is less obvious: while both are recognized in the ASAP analysis, intraspecific distance is below the overall threshold (Fig. 5). However, the ASAP algorithm is more sophisticated than simple distance comparison, and together with other evidence [31], our cox1 data support that these are two species.
Within M. pseudopygmaeus, there are two genetically diverged subgroups that may be recognized as independent partitions with the less-robust ASAP scores (3.5; 5.0) (Fig. 4). One corresponds to haplotypes D, E and F in the haplotype network (Fig. 6) and is likely intraspecific and geography-driven (see below). The second cluster corresponds to specimens from C. affinis; it is further discussed in section “Naticoidea (Caenogastropoda, Littorinimorpha)”. As for all the other M. pseudopygmaeus specimens, their integrity does not seem to be questioned by the cox1 data.
Below, we first summarize all data on the range of first intermediate hosts used by M. pseudopygmaeus, then put these data in the broader digenean context, discuss the likely evolutionary patterns in M. pseudopygmaeus and consider the relevance of our results for evolutionary parasitology.
Gastropod first intermediate hosts of M. pseudopygmaeus: an overview
Littorinoidea (Caenogastropoda, Littorinimorpha)
Periwinkles are the original and the most abundant intertidal first intermediate hosts of M. pseudopygmaeus [31]. Our dataset includes samples from L. obtusata, L. saxatilis, L. arcana (all Atlantic) and L. sitkana (Pacific) and from one more littorinid Lacuna vincta. Patterns of genetic variation among these samples seem to be driven by geography rather than the host species: diverged haplotype E from L. sitkana has North Pacific origin (Fig. 6b). This is further discussed in section “Biogeographic history”.
Littorina scutulata A. Gould, 1849 was also reported to be used by M. pseudopygmaeus as a host [25]. Although adults resulting from the experimental infection of mice with metacercariae isolated from L. scutulata were initially identified as M. pygmaeus [11], revision of the whole mounts clarified that the worms actually were M. pseudopygmaeus [24].
The species M. pseudopygmaeus has not been found in the common periwinkles Littorina littorea (Linnaeus, 1758). Apropos, M. pygmaeus does infect this mollusk, with prevalence as high as 16.7% on the White Sea (unpublished data by D. Fedorov). Neither M. pseudopygmaeus nor other “pygmaeus” microphallids have been found in Littorina squalida Broderip and G. B. Sowerby I, 1829, the Far East sister species of L. littorea (KG, personal observations).
Rissooidea (Caenogastropoda, Littorinimorpha)
Previously, the experimental infections of the common eider with the microphallid metacercariae from O. aculeus produced the maritae identified as M. pseudopygmaeus [29], and the specimen from O. aculeus was identical in 28S rDNA sequence with the specimen of M. pseudopygmaeus from L. saxatilis [31]. Our cox1 data go in line with these findings.
Additionally, M. pseudopygmaeus was reported from another species of Rissooidea, Boreocingula martyni, in the Sea of Okhotsk [31], but we have no molecular evidence to verify this.
Naticoidea (Caenogastropoda, Littorinimorpha)
Metacercariae from the moonsnails C. affinis have previously been identified as M. pseudopygmaeus, despite certain morphological differences [28, 29]. We now found genetically that they are also relatively diverged from the other North Atlantic specimens, with one substitution in ITS2, distant cox1 haplotypes that form a clade on the tree and share a diagnostic combination of three nucleotides. Still, for now we consider the hypothesis that C. affinis is indeed one of the hosts of M. pseudopygmaeus more plausible. This may be a dynamic situation which should be further investigated.
Prevalence of M. pseudopygmaeus infection in C. affinis varies substantially. On Kolguyev Island, Pechora Sea, prevalence is as high as 11.1–25% [29, 33]. In the White and Barents Seas, among 297 C. affinis snails dissected in 2018–2021 [48] only one (0.3%) was found to be infected with M. pseudopygmaeus.
Truncatelloidea (Caenogastropoda, Littorinimorpha)
It is molecularly confirmed that M. pseudopygmaeus infects one species from the superfamily Truncatelloidea, Falsicingula athera Bartsch, 1967 [31], but Peringia ulvae (Pennant, 1777) and Ecrobia ventrosa (Montagu, 1803) (also Truncatelloidea) had never been found infected on the White Sea, with 94,758 and 31,956 specimens examined, respectively [54]. There is a sporadic report of M. pseudopygmaeus infection in E. ventrosa from Skerjafjörður, Iceland [74]. This report could indicate the ongoing colonization of new hosts by M. pseudopygmaeus.
Trochoidea (Vetigastropoda, Trochida)
Microphallus pseudopygmaeus has been known to infect snails belonging to two genera of Trochoidea: Solariella (Solariellidae) and Margarites (Margaritidae). The latter hosts have now been confirmed with molecular data: at least we found no evidence to challenge this fact. Our isolates from M. helicinus (five) and M. groenlandicus (six) grouped in a haplotype network (Fig. 6a). This may reflect certain specialization to the molluscs (Vetigastropoda) that are phylogenetically most distant from the other first intermediate hosts of this parasite (Caenogastropoda), which is further discussed in the section “Host-associated divergence”.
Evolutionary pathways in M. pseudopygmaeus
Biogeographic history
The ancestral first intermediate hosts of the “pygmaeus” microphallids were Littorina spp. which formed in the North Pacific and spread to the Atlantic during the Pliocene opening of the Bering Strait [9], along with their parasites [31]. The glaciation at the end of the Pliocene resulted in the rise of the Bering land bridge (“Beringia”), leading to allopatric speciation in both periwinkles [66, 68] and their microphallids in the Atlantic [31]. Indeed, most species from the “pygmaeus” group infecting only periwinkles occur either in the Atlantic (M. pygmaeus, M. piriformes) or in the Pacific (M. calidris, M. kurilensis).
Occurrence of M. pseudopygmaeus across Eurasia, and likely gene flow between the North Atlantic and North Pacific, is possible because this species has extended the range of its first intermediate host species. The primary role of molluscan hosts is reinforced by the short (8–9 days) lifespan of M. pseudopygmaeus maritae [29], making the long-distance transfer of this parasite within the bird definitive host alone impossible. Arctic waterfowl migrate along Eurasian and North American coastlines, and could potentially facilitate such transfer. However, without a continuous distribution of suitable molluscan hosts along these extensive migration routes, their role in connecting parasite populations remains limited. Instead, successful transfer requires a series of localities where both hosts occur and can support the life cycle.
The common eider, a key definitive host of M. pseudopygmaeus, has circumpolar distribution [78], and so do some of its first intermediate hosts, such as Margarites spp., Solariella spp. and C. affinis. Eiders routinely forage for these molluscs at depths of up to 20 m. Consequently, the life cycle of M. pseudopygmaeus can be completed throughout the Arctic coastal regions, enabling the trans-Arctic dispersal of the parasite. This is in contrast to other species of the “pygmaeus” group microphallids, where the only first intermediate hosts are periwinkles which are absent between the Kara Gates and the Chukchi Sea [66, 67, 72, 73] and cannot support “stepwise” connection between the Pacific and Atlantic populations of parasites.
Given the limited material available from the North Pacific, ideas on the relationships between the North Pacific and the North Atlantic M. pseudopygmaeus are preliminary. Assuming the North Pacific origin of the species [31], the samples from the Sea of Okhotsk would represent an ancestral lineage. These samples include two divergent haplotypes, D and E. A plausible, though tentative, interpretation is that the more distinct haplotype E is closer to the ancestral state, while haplotype D, which is relatively similar to the European variants, might reflect a secondary re-entry into the Pacific from the Atlantic during one of the warm Pleistocene interglacials. Likewise, the occurrence of haplotype F in the North Atlantic could result from a relatively recent colonization event, compared to haplotype A. These hypotheses are now speculative and may change with the accumulation of new data.
Host switching
In principle, host-switching requires that the encounter and compatibility filters [14] open. Encounter filter can open when geographical and/or ecological barriers between the potential hosts are affected by environmental perturbations. In the case of M. pseudopygmaeus, this could have happened during the glaciation cycles in late Pliocene–Pleistocene, when both actual and potential hosts concentrated in the glacial refugia [38, 41]. Compatibility filter can open when a parasite adopts a mechanism to deal with defensive systems of potential new hosts [39]. In the case of M. pseudopygmaeus, this must have been quite massive (considering the host range), but not universal, because some common gastropods, e.g. Buccinidae, and even the periwinkles of the subgenus Littorina (L. littorea and L. squalida) were not colonized.
Microphallus pseudopygmaeus is a good illustration of Digenea in general, where, despite strict specificity to the first intermediate hosts, the major mode of evolution was not co-speciation but host-switching [5, 8, 18, 32]. Colonization of new first intermediate hosts occurred through resource tracking [5]. Among the best-studied examples is host-switching from Planorbidae (Heterobranchia) to Potamiomsidae (Caenogastropoda) in the Schistosoma sinensium lineage, following the uplift of the Tibetan Plateau [3, 53, 57, 79]. Brooks and McLennan [10] suggest two models of resource tracing: phylogenetic tracking (a suitable resource is restricted to the host clade) and sequential colonization (a suitable resource is plesiomorphically or convergently widely spread among the potential hosts). The species M. pseudopygmaeus probably follows the second model: many coastal molluscs can serve as first intermediate host for it, but colonization is limited by a compatibility filter.
Host-associated divergence
Within M. pseudopygmaeus, all the currently available isolates from Margarites spp. grouped in the cox1 haplotype network, suggesting certain intraspecific specialization. This may be attributed to selective success of snail infection, a step towards sympatric speciation by assortative survival [15]. An important question is how it is supported if different molluscan hosts of M. pseudopygmaeus occur in sympatry.
Perhaps specialization could be facilitated in those Arctic areas where periwinkles are absent while Margarites spp. are common, and so are the marine anatids. Rapid egg production by M. pseudopygmaeus maritae (within 2–4 days [24, 29]) allows this parasite to circulate locally in such areas, for example, in Franz-Joseph Land [36]. This assumption is also consistent with lack of divergence in M. pseudopygmaeus from O. aculeus and L. vincta (their distribution ranges are virtually the same as that of Littorina spp. [72]). Distinctness of M. pseudopygmaeus from C. affinis is consistent, too, because these mollusks are also Boreal-Arctic. However, with just two samples, there is not much room for discussion. For both Margarites spp. and C. affinis, it is important to highlight that the specimens in our dataset were collected in the same locations as Littorina, Lacuna and Onoba. Margarites spp. and L. vincta even share the very same microhabitats in the kelp forests.
There are examples of intraspecific genetic differentiation in digeneans guided by the second intermediate or definitive hosts [2, 60], and it can readily be explained by host use preferences or ecological specialization. However, genetic structure corresponding to the first intermediate host species has never before been documented. However, a similar situation has recently been described in the cestode Ligula intestinalis (Linnaeus, 1758) Gmelin, 1790 [56], showing genetic differentiation among the different fish second intermediate host species in sympatry, while also sharing a bird definitive host [60]. The authors believe reproductive isolation may be established despite the continued gene flow, and hypothesize incipient speciation through disruptive selection. In M. pseudopygmaeus, can we estimate whether speciation is ongoing?
Possible speciation
Though the hypothesis remains speculative, it is worth discussing that host-associated divergence in M. pseudopygmaeus may be a step towards speciation.
The life cycle of M. pseudopygmaeus involves passive transmission only: the eggs get accidentally ingested by the snails and the snails get ingested by the birds, mostly eiders. So, no behavioural features of a parasite may have contributed to host switching and the potential speciation. Instead, M. pseudopygmaeus miracidium, mother and daughter sporocysts have probably gained characters necessary to successfully establish infection in the new snail hosts. Parthenogenetic reproduction of these successful mutants would result in hundreds (and as many as 3,000) metacercariae sharing this genotype, enhancing the spread of the newly emerged form.
Next, we would expect selection against the intermediate forms – those poor at infecting both the original and the new hosts. Disruptive selection is plausible because we detected divergence in the most unrelated hosts, Margarites spp., which are more likely to bear drastic differences in the defence mechanisms, intestinal environment, haemocoel composition, etc. If the other hosts share more of these features, they can be tackled by the genetically similar parasites, not yielding any specialization. The species C. affinis is also quite distinct in its ecology (shell-boring predators, not grazers) and geographical distribution, and samples from this host are genetically diverged. Due to limited sample size for C. affinis, discussion below focuses mostly on Margarites spp.
For the reproductive isolation to endure, assortative survival in the first intermediate hosts must be followed by assortative mating in the definitive hosts. On the one hand, the eiders’ diet includes all the M. pseudopygmaeus first intermediate hosts. Mating between maritae originating from different gastropod hosts seems unavoidable at first, and this would contribute to the constant genetic mixing in a parasite. On the other, there are plausible mechanisms that could support consistent mating among maritae tracing the same molluscan origin. First, the M. pseudopygmaeus maritae are very short-lived and also tend to pass through the bird’s intestine in a compact yet numerous group originating from the same infected snail, likely mating within this group. Moreover, selfing may be suspected in extremely small M. pseudopygmaeus maritae (around 300 μm long) that invariably contain embryonated eggs in natural infections (KG, unpublished results). The probability of selfing in digeneans is not clear, and in microphallids specifically it has been tested by culturing individual worms in vitro, with both positive [12] and negative [19, 64] results.
Distinct local transmission hotspots in high latitudes (discussed section “Naticoidea (Caenogastropoda, Littorinimorpha)”) and in Pleistocene glacial refugia could also contribute to the possibility of speciation. The definitive hosts, common eiders, concentrate in coastal areas of islands for breeding, and in Franz Josef Land infected Margarites spp. occur precisely in the vicinity of these colonies [36]. Such hotspots are likely to be partially isolated from one another due to the short lifespan of the maritae. This isolation may favor mutation accumulation in local parasite populations associated with local first intermediate hosts.
Conclusion
Taken together, our data confirm the exceptionally wide range of gastropods that serve as first intermediate hosts for the digenean M. pseudopygmaeus. This species also demonstrates the genetic divergence associated with its vetigastropod hosts, Margarites spp., and most probably with its naticid host C. affinis. This makes it an excellent host-parasite system to explore evolution of host specificity, host switching, and speciation. This model could be meaningful even beyond the parasite discourse because specialization within M. pseudopygmaeus is likely reinforced in the Arctic regions, and in circumstances comparable to those that occurred during the Pleistocene glacial cycles when many modern species formed.
Acknowledgments
We are grateful to two anonymous reviewers for their valuable contributions to a better integrity of this manuscript. We thank the field stations where sampling essential for this study took place: the White Sea Biological Station “Kartesh”, Zoological Institute (ZIN RAS); the Educational and Research Station “Belomorskaia”, St Petersburg University (SPbU); the White Sea Biological Station, Moscow State University; and the Seasonal Biological Station “Dalniye Zelentsy”, Murmansk Marine Biological Institute. Valuable samples from the Pechora Sea were accessible thanks to the operation of the RV “Professor Vladimir Kuznetsov” (ZIN RAS). Special thanks are due to Dr. K.V. Regel (Institute of Biological Problems of the North, Magadan, Russia) and Prof. K. Skírnisson (University of Iceland, Reykjavik, Iceland) for their help during the fieldwork in the Sea of Okhotsk and Iceland, respectively. We appreciate excellent sequencing performed for us by Dr. Anna Romanovitch at the Resource centre “Molecular and Cell Technologies” of the Research Park, SPbU. This work was supported by the Russian Science Foundation (23-14-00329), and sampling at the White Sea was supported by the State Academic Program, Russia (125012800903-5).
Conflicts of interest
The authors declare that they have no conflict of interest.
Supplementary material
Supplementary Table S1: The average pairwise genetic distances in 28S rDNA fragment (971 bp).
Supplementary Table S2: The average pairwise genetic distances in ITS2–5.8S rDNA fragment (266 bp).
Supplementary Table S3: p-distances for sequences from specimens of Microphallus of the „pygmaeus“ group and M. similis.
Supplementary File: ASAP_samples_104_801.
Access Supplementary MaterialReferences
- Achatz TJ, Pulis EE, Junker K, Binh TT, Snyder SD, Tkach VV. 2019. Molecular phylogeny of the Cyathocotylidae (Digenea, Diplostomoidea) necessitates systematic changes and reveals a history of host and environment switches. Zoologica Scripta, 48, 545–556. [Google Scholar]
- Agustina V, Saichua P, Laha T, Tangkawatana S, Prakobwong S, Laoprom N, Kamphasri W, Chareonchai C, Blair D, Suttiprapa S. 2024. Exploring the second intermediate hosts and morphology of human- and cat-specific Opisthorchis viverrini-like populations. International Journal for Parasitology, 54, 497–506. [Google Scholar]
- Attwood SW, Upatham ES, Meng XH, Qiu D-C, Southgate VR. 2002. The phylogeography of Asian Schistosoma (Trematoda: Schistosomatidae). Parasitology, 125, 99–112. [CrossRef] [PubMed] [Google Scholar]
- Belopolskaja MM. 1963. Family Microphallidae Travassos, 1920. Osnovy Trematodologii, 21, 259–502. [Google Scholar]
- Blair D, Davis GM, Wu B. 2001. Evolutionary relationships between trematodes and snails emphasizing schistosomes and paragonimids. Parasitology, 123, 229–243. [Google Scholar]
- Bowles J, Blair D, McManus DP. 1995. A molecular phylogeny of the human schistosomes. Molecular Phylogenetics and Evolution, 4, 103–109. [CrossRef] [PubMed] [Google Scholar]
- Bowles J, Hope M, Tiu WU, Liu X, McManus DP. 1993. Nuclear and mitochondrial genetic markers highly conserved between Chinese and Philippine Schistosoma japonicum. Acta Tropica, 55, 217–229. [PubMed] [Google Scholar]
- Brant SV, Loker ES. 2013. Discovery-based studies of schistosome diversity stimulate new hypotheses about parasite biology. Trends in Parasitology, 29, 449–459. [Google Scholar]
- Briggs JC. 2003. Guest editorial: Marine centres of origin as evolutionary engines. Journal of Biogeography, 30, 1–18. [Google Scholar]
- Brooks DR, McLennan DA. 1993. Parascript – Parasites and the language of evolution. Washington: Smithsonian Institution Press. [Google Scholar]
- Ching HL. 1961. Three trematodes from the harlequin duck. Canadian Journal of Zoology, 39, 373–376. [Google Scholar]
- Chuang Y, Shinn AP, Bron JE. 2025. In vitro culture and confocal microscopy study of Maritrema gratiosum Nicoll, 1907 (Digenea): From metacercaria to ovigerous adult. Parasitology Research, 124, 1–17. [Google Scholar]
- Clement M, Snell Q, Walke P, Posada D, Crandall K. 2002. TCS: estimating gene genealogies, in Proceedings of the 16th International Parallel and Distributed Processing Symposium. Ft. Lauderdale, FL: IEEE. p. 7. [Google Scholar]
- Combes C. 2001. Parasitism: The ecology and evolution of intimate interactions. Chicago, IL: University of Chicago Press. [Google Scholar]
- Combes C, Théron A. 2000. Metazoan parasites and resource heterogeneity: Constraints and benefits. International Journal for Parasitology, 30, 299–304. [Google Scholar]
- Corel Corporation. 2022. CorelDRAW Graphics Suite [computer program]. Version 24.0.0.301. Ottawa, Canada. [Google Scholar]
- Cribb TH, Anderson GR, Adlard RD, Bray RA. 1998. A DNA-based demonstration of a three-host life-cycle for the Bivesiculidae (Platyhelminthes: Digenea). International Journal for Parasitology, 28, 1791–1795. [PubMed] [Google Scholar]
- Cribb TH, Bray RA, Olson PD, Littlewood DTJ. 2003. Life cycle evolution in the Digenea: A new perspective from phylogeny, in The Evolution of Parasitism – A Phylogenetic Perspective, Littlewood DTJ, Editor. Academic Press: London. p. 197. [Google Scholar]
- Daczkowski CM, Pung OJ. 2016. In vitro insemination of the microphallid digenean Gynaecotyla adunca. Journal of Parasitology, 102, 5–10. [Google Scholar]
- Edler D, Klein J, Antonelli A, Silvestro D. 2021. raxmlGUI 2.0: A graphical interface and toolkit for phylogenetic analyses using RAxML. Methods in Ecology and Evolution, 12, 373–377. [Google Scholar]
- Fedosov A, Achaz G, Gontchar A, Puillandre N. 2022. mold, a novel software to compile accurate and reliable DNA diagnoses for taxonomic descriptions, Molecular Ecology Resources, 22, 2038–2053. [Google Scholar]
- Fraija-Fernández N, Olson PD, Crespo EA, Raga JA, Aznar FJ, Fernández M. 2015. Independent host switching events by digenean parasites of cetaceans inferred from ribosomal DNA. International Journal for Parasitology, 45, 167–173. [Google Scholar]
- Gagnon DK, Detwiler JT. 2019. Broader geographic sampling increases extent of intermediate host specificity for a trematode parasite (Notocotylidae: Quinqueserialis quinqueserialis). Journal of Parasitology, 105, 874. [Google Scholar]
- Galaktionov KV. 2009. Description of maritae and determination of the species status of Microphallus pseudopygmaeus sp. nov. (Trematoda: Microphallidae). Parazitologiya, 43, 289–299. [Google Scholar]
- Galaktionov KV. 1980. Four types of metacercariae species in molluscs, Littorina saxatilis and L. obtusata from the Barents and White Seas. Vestnik Leningradskogo Universiteta, 3, 21–28. [Google Scholar]
- Galaktionov KV. 1983. Microphallids in the pygmaeus group. 1. Description of the species Microphallus pygmaeus (Levinsen 1881) nec Odhner, 1905 and M. piriformes (Odhner, 1905) nom. nov. (Trematoda, Microphallidae). Vestnik Leningradskogo Universiteta Biologiya, 1983(3), 20–30. [Google Scholar]
- Galaktionov KV. 1984. Microphallids of the “pygmaeus” group II. Microphallus triangulatus sp. nov. (Trematoda: Microphallidae). Vestnik Leningradskogo Universiteta, 3, 5–11. [Google Scholar]
- Galaktionov KV. 1986. Metacercariae from the family Microphallidae Travassos, 1920 in benthic mollusks in the south-eastern part of the Barents Sea. Parazitologiya, 20, 389–396. [Google Scholar]
- Galaktionov KV. 1993. Life cycles of trematodes as components of ecosystems. Apatity: Kola Scientific Centre of the Russian Academy of Sciences. [Google Scholar]
- Galaktionov KV, Blasco-Costa I. 2018. Microphallus ochotensis sp. nov. (Digenea, Microphallidae) and relative merits of two-host microphallid life cycles. Parasitology Research, 117, 1051–1068. [Google Scholar]
- Galaktionov KV, Blasco-Costa I, Olson PD. 2012. Life cycles, molecular phylogeny and historical biogeography of the “pygmaeus” microphallids (Digenea: Microphallidae): Widespread parasites of marine and coastal birds in the Holarctic. Parasitology, 139, 1346–1360. [Google Scholar]
- Galaktionov KV, Dobrovolskij AA. 2003. The trematode life cycle as a system of adaptations, in The biology and evolution of trematodes: An essay on the biology, morphology, life cycles, transmissions, and evolution of digenetic trematodes. Springer Netherlands: Dordrecht. [Google Scholar]
- Galaktionov KV, Marasaev SF. 1986. Ecological analysis of the trematode fauna of benthic mollusks in the south-eastern zone of the Barents Sea. Parazitologiya, 20, 455–460. [Google Scholar]
- Galaktionov KV, Regel KV, Atrashkevich GI. 2010. Microphallus kurilensis sp. nov., a new species of microphallids from the pygmaeus species group (Trematoda, Microphallidae) from the coastal areas of Okhotsk and Bering Seas. Parazitologiia, 44, 496–507. [Google Scholar]
- Galaktionov KV, Solovyeva AI, Miroliubov A. 2021. Elucidation of Himasthla leptosoma (Creplin, 1829) Dietz, 1909 (Digenea, Himasthlidae) life cycle with insights into species composition of the north Atlantic Himasthla associated with periwinkles Littorina spp. Parasitology Research, 120, 1649–1668. [Google Scholar]
- Galaktionov KV, Węsławski JM, Stempniewicz L. 2021. Food chain, parasites and climate changes in the high Arctic: A case study on trophically transmitted parasites of common eider Somateria mollissima at Franz Josef Land. Polar Biology, 44, 1321–1342. [Google Scholar]
- Gonchar A, Galaktionov KV. 2021. It is marine: distinguishing a new species of Catatropis (Digenea: Notocotylidae) from its freshwater twin. Parasitology, 148, 74–83. [Google Scholar]
- Hoberg EP. 1992. Congruent and synchronic patterns in biogeography and speciation among seabirds, pinnipeds, and cestodes. Journal of Parasitology, 78, 601. [Google Scholar]
- Hoberg EP, Brooks DR. 2008. A macroevolutionary mosaic: Episodic host-switching, geographical colonization and diversification in complex host–parasite systems. Journal of Biogeography, 35, 1533–1550. [Google Scholar]
- Hoberg EP, Brooks DR. 2015. Evolution in action: Climate change, biodiversity dynamics and emerging infectious disease. Philosophical Transactions of the Royal Society B, 370, 20130553. [Google Scholar]
- Hoberg EP, Cook JA, Agosta SJ, Boeger W, Galbreath KE, Laaksonen S, Kutz SJ, Brooks DR. 2017. Arctic systems in the Quaternary: Ecological collision, faunal mosaics and the consequences of a wobbling climate. Journal of Helminthology, 91, 409–421. [Google Scholar]
- Jägerskiöld LA. 1900. Levinsenia (Distomum) pygmaea Levinsen, ein genitalnapftragendes Distomum. Zentralblatt für Bakteriologie, 1, 732–740. [Google Scholar]
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A. 2012. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics, 28, 1647–1649. [CrossRef] [PubMed] [Google Scholar]
- Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A. 2019. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics, 35, 4453–4455. [CrossRef] [PubMed] [Google Scholar]
- Králová-Hromadová I, Špakulová M, Horáčková E, Turčeková L, Novobilský A, Beck R, Koudela B, Marinculić A, Rajský D, Pybus M. 2008. Sequence analysis of ribosomal and mitochondrial genes of the giant liver fluke Fascioloides magna (Trematoda: Fasciolidae): Intraspecific variation and differentiation from Fasciola hepatica. Journal of Parasitology, 94, 58–67. [Google Scholar]
- Kremnev G, Gonchar A, Krapivin V, Knyazeva O, Krupenko D. 2020. First elucidation of the life cycle in the family Brachycladiidae (Digenea), parasites of marine mammals. International Journal for Parasitology, 50, 997–1009. [Google Scholar]
- Kremnev G, Gonchar A, Krapivin V, Uryadova A, Miroliubov A, Krupenko D. 2021. Life cycle truncation in Digenea, a case study of Neophasis spp. (Acanthocolpidae), International Journal for Parasitology: Parasites and Wildlife, 15, 158–172. [Google Scholar]
- Kremnev G, Gonchar A, Uryadova A, Krapivin V, Skobkina O, Gubler A, Krupenko D. 2023. No tail no fail: Life cycles of the Zoogonidae (Digenea). Diversity, 15, 121. [Google Scholar]
- Kremnev G, Krapivin V, Krupenko D. 2025. Second intermediate hosts range and their infection patterns in Orthosplanchnus arcticus, a potentially zoonotic parasite from the family Brachycladiidae (Digenea). Journal of Helminthology, 99, 109. [Google Scholar]
- Krupenko D, Kremnev G, Gonchar A, Gubler A, Skobkina O. 2024. Wandering the taxonomic mine-field: the Podocotyle species complex (Digenea: Opecoelidae). Systematic Parasitology, 101, 72. [Google Scholar]
- Krupenko D, Kremnev G, Gonchar A, Uryadova A, Miroliubov A, Krapivin V, Skobkina O, Gubler A, Knyazeva O. 2022. Species complexes and life cycles of digenetic trematodes from the family Derogenidae. Parasitology, 149, 1590–1606. [CrossRef] [PubMed] [Google Scholar]
- Laidemitt MR, Brant SV, Mutuku MW, Mkoji GM, Loker ES. 2019. The diverse echinostomes from East Africa: With a focus on species that use Biomphalaria and Bulinus as intermediate hosts. Acta Tropica, 193, 38–49. [Google Scholar]
- Lawton SP, Hirai H, Ironside JE, Johnston DA, Rollinson D. 2011. Genomes and geography: Genomic insights into the evolution and phylogeography of the genus Schistosoma. Parasites & Vectors, 4, 131. [Google Scholar]
- Levakin IA, Nikolaev KE, Galaktionov KV. 2013. Long-term variation in trematode (Trematoda, Digenea) component communities associated with intertidal gastropods is linked to abundance of final hosts. Hydrobiologia, 706, 103–118. [Google Scholar]
- Levinsen GMR. 1881. Bidrag til kundskab om Grønlands trematodfauna. Oversigt over det Kongelige Danske Videnskabernes Selskabs Forhandlinger. 52, 84. [Google Scholar]
- Linnaeus C. 1758. Taenia intestinalis, in Systema naturae per regna tria naturae, 10th edn, vol. 1, Holmiae (Stockholm): Laurentii Salvii. p. 824. [Google Scholar]
- Lockyer AE, Olson PD, Østergaard P, Rollinson D, Johnston DA, Attwood SW, Southgate VR, Horak P, Snyder SD, Le TH, Agatsuma T, McManus DP, Carmichael AC, Naem S, Littlewood DTJ. 2003. The phylogeny of the Schistosomatidae based on three genes with emphasis on the interrelationships of Schistosoma Weinland, 1858. Parasitology, 126, 203–224. [CrossRef] [PubMed] [Google Scholar]
- Louvard C, Corner RD, Cutmore SC, Cribb TH. 2022. Evidence that host ecology drives first intermediate host use in the Didymozoidae (Trematoda: Hemiuroidea): An asexual infection in a vermetid (Gastropoda). Journal of Helminthology, 96(e88), 1–11. [CrossRef] [Google Scholar]
- Martin SB, Sasal P, Cutmore SC, Ward S, Aeby GS, Cribb TH. 2018. Intermediate host switches drive diversification among the largest trematode family: evidence from the Polypipapiliotrematinae n. subf. (Opecoelidae), parasites transmitted to butterflyfishes via predation of coral polyps. International Journal for Parasitology, 48, 1107–1126. [Google Scholar]
- Nazarizadeh M, Nováková M, Drábková M, Catchen J, Olson PD, Štefka J. 2024. Highly resolved genome assembly and comparative transcriptome profiling reveal genes related to developmental stages of tapeworm Ligula intestinalis. Proceedings of the Royal Society B, 291, 20232563. [Google Scholar]
- Pakharukova MY, Mordvinov VA. 2022. Similarities and differences among the Opisthorchiidae liver flukes: insights from Opisthorchis felineus. Parasitology, 149, 1306–1318. [Google Scholar]
- Posit Team. 2025. RStudio: Integrated Development Environment for R. Posit Software. Boston, MA: PBC. [Google Scholar]
- Puillandre N, Brouillet S, Achaz G. 2021. ASAP: Assemble species by automatic partitioning. Molecular Ecology Resources, 21, 609–620. [CrossRef] [PubMed] [Google Scholar]
- Pung OJ, Lester T, Burger AR, Alyanak E, O’Leary PA. 2011. Optimization of culture conditions for in vitro fertilization and reproduction of Microphallus turgidus (Trematoda: Microphallidae). Journal of Parasitology, 97, 1–7. [Google Scholar]
- R Core Team. 2016. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. [Google Scholar]
- Reid DG. 1996. Systematics and evolution of Littorina. London: The Ray Society. [Google Scholar]
- Reid DG. 2001. New data on the taxonomy and distribution of the genus Littoraria Griffith and Pidgeon, 1834 (Gastropoda: Littorinidae) in Indo-West Pacific mangrove forests. Nautilus, 115, 115–139. [Google Scholar]
- Reid DG, Dyal P, Williams ST. 2012. A global molecular phylogeny of 147 periwinkle species (Gastropoda, Littorininae). Zoologica Scripta, 41, 125–136. [Google Scholar]
- Repkin EA, Maltseva AL, Varfolomeeva MA, Aianka RV, Mikhailova NA, Granovitch AI. 2020. Genetic and morphological variation of metacercariae of Microphallus piriformes (Trematoda, Microphallidae): Effects of paraxenia and geographic location. International Journal for Parasitology: Parasites and Wildlife, 11, 235–245. [Google Scholar]
- Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. 2012. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61, 539–542. [CrossRef] [PubMed] [Google Scholar]
- Saville DH, Galaktionov KV, Irwin SWB, Malkova II. 1997. Morphological comparison and identification of metacercariae in the “pygmaeus” group of microphallids, parasites of seabirds in western palaearctic regions. Journal of Helminthology, 71, 167–174. [Google Scholar]
- Sirenko BI. 2001. List of species of free-living invertebrates of Eurasian Arctic seas and adjacent deep waters. St. Petersburg: Zoological Institute, Russian Academy of Sciences. [Google Scholar]
- Sirenko BI. 2013. Check-list of species of free-living invertebrates of the Russian Far Eastern seas. St. Petersburg: Zoological Institute, Russian Academy of Sciences. [Google Scholar]
- Skírnisson K, Galaktionov KV. 2002. Life cycles and transmission patterns of seabird digeneans in SW Iceland. Sarsia, 87, 144–151. [Google Scholar]
- Tamura K, Stecher G, Kumar S. 2021. MEGA11: Molecular Evolutionary Genetics Analysis version 11. Molecular Biology and Evolution, 38, 3022–3027. [CrossRef] [PubMed] [Google Scholar]
- Tkach VV, Littlewood DTJ, Olson PD, Kinsella JM, Swiderski Z. 2003. Molecular phylogenetic analysis of the Microphalloidea Ward, 1901 (Trematoda: Digenea). Systematic Parasitology, 56, 1–15. [PubMed] [Google Scholar]
- Vences M, Miralles A, Brouillet S, Ducasse J, Fedosov A, Kharchev V, Kumari S, Patmanidis S, Puillandre N, Scherz MD, Kostadinov I, Renner SS. 2021. iTaxoTools 0.1: Kickstarting a specimen-based software toolkit for taxonomists. Megataxa, 6, 77–92. [Google Scholar]
- Waltho C, Coulson J. 2015. The Common Eider. London: T. & A. D. Poyser. [Google Scholar]
- Webster BL, Littlewood DTJ. 2012. Mitochondrial gene order change in Schistosoma (Platyhelminthes: Digenea: Schistosomatidae). International Journal for Parasitology, 42, 313–321. [Google Scholar]
- Wee NQ, Cribb TH, Corner RD, Ward S, Cutmore SC. 2021. Gastropod first intermediate hosts for two species of Monorchiidae Odhner, 1911 (Trematoda): I can’t believe it’s not bivalves! International Journal for Parasitology, 51, 1035–1046. [Google Scholar]
- Wickham H. 2016. ggplot2: Elegant Graphics for Data Analysis. New York: Springer-Verlag. [Google Scholar]
Cite this article as: : Sokolova AI, Galaktionov KV & Gonchar A. 2026. Microphallus pseudopygmaeus (Digenea) infects phylogenetically distant gastropods, with signs of host-linked genetic divergence. Parasite 33, 55. https://doi.org/10.1051/parasite/2026016.
All Tables
All Figures
 |
Figure 1 Metacercariae of microphallids of the “pygmaeus” group, microphotographs. (a) M. pseudopygmaeus, (b) M. pygmaeus, (c) M. triangulatus, (d) M. kurilensis, (e) M. piriformes, (f) M. calidris. |
| In the text | |
 |
Figure 2 Life cycle of microphallids of the “pygmaeus” group. Definitive hosts: marine birds; first intermediate hosts: marine gastropods (typically, periwinkles); trophic transmission: the Pac-Man icon. |
| In the text | |
 |
Figure 3 Map of all sampling sites (indicated by arrows) and M. pseudopygmaeus sampling sites (red dots). Closely spaced localities (e.g., within the Pechora Sea and the White Sea) are not shown on the map. |
| In the text | |
 |
Figure 4 Phylogenetic relationships of the “pygmaeus” group microphallids based on cox1 sequences. (a) Bayesian phylogenetic tree. Poorly supported nodes (posterior probability < 0.75) were collapsed into polytomies. For nodes also recovered in the maximum likelihood (ML) analysis, both BI and ML support values are shown. (b) Five best ASAP scores and partition numbers (in brackets); and color-coded putative species boundaries corresponding to the clades in the tree. |
| In the text | |
 |
Figure 5 Frequency distribution of pairwise genetic distances in the cox1 gene fragment (803 bp) for seven species of Microphallus. MPG, microphallids of the “pygmaeus” group. |
| In the text | |
 |
Figure 6 Haplotype networks for M. pseudopygmaeus, also including several samples of M. pygmaeus and M. similis, based on the fragment of the cox1 gene (800 bp). Circle sizes represent haplotype frequencies, hatch marks represent substitutions (if more than five, replaced with numbers). The asterisk marks the sample from L. arcana (belongs to “saxatilis” species complex). Haplotypes are colored according to parasite host species (a) and sampling location (b). The sampling locations in the Barents Sea: Dalniye Zelentsy (southwestern), Chosha Bay, Kanin Cape (southeastern); in the Pechora Sea: Dolgiy Island, Vaygach Island, Malyi Zincovyi Island, Gubistyi Island; in the White Sea: Levin Navolok Bay, Cape Kindo, Korovya Varakka Cliff; in the Sea of Okhotsk: Tyuleniy Island, Taygonos Cape, Nagaev Bay; in the Atlantic Ocean: Grindavik. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


