| Issue |
Parasite
Volume 32, 2025
|
|
|---|---|---|
| Article Number | 63 | |
| Number of page(s) | 10 | |
| DOI | https://doi.org/10.1051/parasite/2025056 | |
| Published online | 25 September 2025 | |
Research Article
Cryptosporidium spp. in wild rodents in the Poyang Lake region, China
Cryptosporidium spp. chez les rongeurs sauvages de la région du lac Poyang, en Chine
1
National Institute of Parasitic Diseases, Chinese Center for Disease Control and Prevention (Chinese Center for Tropical Diseases Research), NHC Key Laboratory of Parasite and Vector Biology, National Key Laboratory of Intelligent Tracking and Forecasting for Infectious Diseases, WHO Collaborating Centre for Tropical Diseases, National Center for International Research on Tropical Diseases, Shanghai 200025, PR China
2
Institute of Biological Resources, Jiangxi Academy of Sciences, Nanchang 330096, PR China
3
The School of Global Health, Chinese Center for Tropical Diseases Research, Shanghai Jiao Tong University School of Medicine, Shanghai 200025, PR China
* Corresponding authors: This email address is being protected from spambots. You need JavaScript enabled to view it.
(Yujuan Shen); This email address is being protected from spambots. You need JavaScript enabled to view it.
(Jianping Cao)
Received:
17
April
2025
Accepted:
6
September
2025
Abstract
Cryptosporidium spp. are zoonotic protozoan parasites that cause diarrheal disease worldwide. Rodents can harbor diverse Cryptosporidium spp. and facilitate their transmission to the environment and other hosts, including humans. However, data on Cryptosporidium infection in wild rodents in the Poyang Lake region, China’s largest freshwater lake, remain scarce. Here, we investigated Cryptosporidium spp. in 273 wild rodents collected from seven sites adjacent to villages around Poyang Lake between 2022 and 2024. The rodents were identified by cytochrome c oxidase subunit I (COI) gene sequencing as Apodemus agrarius (n = 148) and Rattus losea (n = 125). Nested PCR targeting the small subunit ribosomal RNA (SSU rRNA) gene revealed an overall Cryptosporidium spp. infection rate of 16.5% (45/273, 95% CI: 12.3–21.9%), with 20.3% (30/148, 95% CI: 14.2–27.8%) in A. agrarius and 12.0% (15/125, 95% CI: 6.9–19.0%) in R. losea. Sequence and phylogenetic analyses identified seven Cryptosporidium species/genotypes: C. apodemi, C. canis, C. muris, C. suis, C. ubiquitum, rat genotype II, and rat genotype III. Notably, the detection of four zoonotic species (C. canis, C. muris, C. suis, and C. ubiquitum) highlights the potential risk of zoonotic transmission of Cryptosporidium spp. from wild rodents to humans in this region. These findings underscore the need for systematic surveillance and control of Cryptosporidium spp. in wild rodent communities around Poyang Lake.
Résumé
Les Cryptosporidium spp. sont des parasites protozoaires zoonotiques responsables de maladies diarrhéiques dans le monde entier. Les rongeurs peuvent héberger diverses espèces de Cryptosporidium et faciliter leur transmission à l’environnement et à d’autres hôtes, dont l’homme. Cependant, les données sur l’infection à Cryptosporidium chez les rongeurs sauvages de la région du lac Poyang, le plus grand lac d’eau douce de Chine, restent rares. Nous avons étudié Cryptosporidium spp. chez 273 rongeurs sauvages collectés dans sept sites adjacents à des villages autour du lac Poyang entre 2022 et 2024. Les rongeurs ont été identifiés par séquençage du gène de la sous-unité I de la cytochrome c oxydase (COI) comme étant Apodemus agrarius (n = 148) et Rattus losea (n = 125). La PCR imbriquée ciblant le gène de l’ARN ribosomique de la petite sous-unité (ARNr SSU) a révélé un taux d’infection global par Cryptosporidium spp. de 16,5 % (45/273, IC à 95 % : 12,3-21,9 %), avec 20,3 % (30/148, IC à 95 % : 14,2-27,8 %) chez A. agrarius et 12,0 % (15/125, IC à 95 % : 6,9-19,0 %) chez R. losea. Les analyses de séquence et phylogénétiques ont identifié sept espèces/génotypes de Cryptosporidium : C. apodemi, C. canis, C. muris, C. suis, C. ubiquitum, génotype II du rat, génotype III du rat. Notamment, la détection de quatre espèces zoonotiques (C. canis, C. muris, C. suis et C. ubiquitum) met en évidence le risque potentiel de transmission zoonotique de Cryptosporidium spp. des rongeurs sauvages à l’homme dans cette région. Ces résultats soulignent la nécessité d’une surveillance et d’un contrôle systématiques de Cryptosporidium spp. dans les communautés de rongeurs sauvages autour du lac Poyang.
Key words: Cryptosporidium / Zoonotic / Wild rodents / Apodemus agrarius / Rattus losea / Poyang Lake region
Edited by: Jean-Lou Justine
These authors contributed equally.
© Q. Zuo et al., published by EDP Sciences, 2025
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Introduction
Cryptosporidiosis, a zoonotic enteric disease caused by Cryptosporidium spp. parasites, poses a major health risk to humans and animals worldwide [10]. Infections may occur through ingestion of Cryptosporidium oocysts via contaminated water and food, or direct contact with infected hosts [11]. This disease can cause severe and even life-threatening diarrheal illness in vulnerable populations, particularly in children and immunocompromised individuals [35]. To date, among the more than 170 species/genotypes of Cryptosporidium reported in various animals, 23 species and two genotypes are known to infect humans [12]. Understanding of the transmission dynamics and potential risks of Cryptosporidium spp. in populations in close contact with humans is key to developing effective prevention and control strategies.
Rodents, the most abundant mammals, are known for their adaptability and resilience, particularly because of their wide distribution near human settlements [3]. Many rodents are highly contagious carriers of zoonotic diseases, such as plague [1], hantavirus infection [36], and cryptosporidiosis [12]. To date, rodents have been confirmed to have been infected with more than 26 Cryptosporidium species and 59 genotypes. In particular, C. hominis, C. parvum, C. meleagridis, C. felis, C. viatorum, C. mortiferum, C. muris, C. andersoni, C. suis, C. scrofarum, C. equi, C. erinacei, C. tyzzeri, C. occultus, C. ditrichi, C. wrairi, and the skunk genotype are known to infect humans [12]. These diverse pathogens in rodents might pose a major zoonotic disease threat to the environment and to human populations.
A recent study has highlighted a substantial burden of human cryptosporidiosis in Jiangxi Province, where the total number of cases accounted for 13.95% (694/4975) of the national total, ranking second nationwide [43]. Furthermore, Cryptosporidium spp. infections have been documented in various farmed and companion animals, including bamboo rats (Rhizomys sinensis, 33.3%, 51/153) [23], cattle (Bos taurus, 12.8%, 71/556) [26], peafowls (Pavo cristatus, 9.52%, 10/105) [17], pigs (Sus scrofa domesticus, 4.0%, 41/1036) [42], and masked palm civets (Paguma larvata, 1.0%, 5/489) [46]. However, data on Cryptosporidium infections in wild animals remain relatively limited.
Poyang Lake (115°49′E–116°46′E, 28°24′N–29°46′N), the largest freshwater lake in China, is a critical source of drinking water for Jiangxi and neighboring provinces [20]. In recent years, an extensive and dense rodent community has been observed in the Poyang Lake region, which has been reported to have potential for a future rodent population outbreak [6]. These rodents are not only abundant in the lake region but also commonly found in agricultural areas and human settlements [3], thus posing potential public health concerns in sylvatic environments. The expansion of rodent populations might facilitate the transmission of pathogens, particularly waterborne pathogens. Therefore, investigating the presence and dynamics of Cryptosporidium spp. pathogens in the wild rodent communities within Poyang Lake’s ecosystem is crucial to safeguard public health, preserve ecological balance, and ensure sustainable development.
In this study, we aimed to investigate the infection status, genetic characteristics, and zoonotic potential of Cryptosporidium spp. in the wild rodent communities in the Poyang Lake region.
Materials and methods
Study area
Field studies were conducted around Poyang Lake (28.24°N–29.46°N, 115.49°E–116.75°E), the largest freshwater lake in China, located in northern Jiangxi Province along the middle and lower reaches of the Yangtze River. In this region, the subtropical monsoon climate, with an annual average temperature of 17 °C to 18 °C, and the abundance of herbaceous vegetation, dominated by Carex spp. and Phragmites spp., provide suitable habitats for wild rodents, such as the striped field mouse (Apodemus agrarius) and the losea rat (Rattus losea). We selected seven sampling sites in farmlands and lakeside areas adjacent to both villages and water sources for wild rodent collection (Fig. 1 and Table 1).
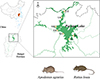 |
Figure 1 Study area and sampling sites (marked with yellow squares) around Poyang Lake, Jiangxi Province, China. |
Information on Cryptosporidium spp. in wild rodents in the Poyang Lake region.
Sample collection and preparation
This study involved 273 fecal samples from wild rodents, which were collected at seven sampling sites from 2022 to 2024 (Fig. 1, Table 1). Rodents were captured with snap traps (150 × 80 mm, Guixi Mousing Tool Factory, Jiangxi, China) baited with fresh sunflower seeds and then transported to the laboratory.
The sex of the rodents was determined through morphological observation of the genitalia. The rodents’ age was estimated according to body weight and used to divide the rodents into groups [19, 24]. Apodemus agrarius were divided into a juvenile group (body weight ≤ 16.0 g), sub-adult group (body weight = 16.1–23.0 g), adult group (body weight = 23.1–37.0 g), and old group (body weight > 37.0 g). Rattus losea were divided into a juvenile group (body weight ≤ 37.0 g), sub-adult group (body weight = 37.1–65.0 g), adult group (body weight = 65.1–92.0 g), and old group (body weight > 92.0 g). Fecal samples were placed in labeled 50 mL centrifuge tubes and stored separately at −20 °C before DNA extraction.
The protocol was approved by the Animal Experiment Ethics Committee of the Institute of Biological Resources, Jiangxi Academy of Sciences (No. 2022023).
DNA extraction
Genomic DNA from ~200 mg rodent feces was extracted with a QIAamp DNA Stool Mini kit (Cat. #51604; QIAGEN, Hilden, Germany) according to the manufacturer’s instructions. The DNA was stored at −20 °C until PCR analysis.
Host identification
A 750 bp fragment of the mtDNA cytochrome oxidase I (COI) gene was amplified with the primers BatL5310 and R6036R for rodent species identification. PCR conditions and primer design were as described by Robins et al. [33]. Each PCR reaction comprised 35 cycles of denaturation at 94 °C for 45 s, annealing at 54 °C for 60 s, and extension at 72 °C for 60 s. All amplified PCR products were visualized through 1% agarose gel electrophoresis. All successful amplifications were sent to Sangon Biotech (Shanghai) Co., Ltd. (Shanghai, China) for sequencing and then compared with entries in the NCBI database (http://www.ncbi.nlm.nih.gov/BLAST) for rodent species identification.
Pathogen identification
Cryptosporidium species were determined with nested PCR and sequence analysis of the small subunit (SSU) rRNA gene [44]. Subsequently, the 60 kDa glycoprotein (gp60) gene was amplified with nested PCR to further subtype the detected zoonotic Cryptosporidium spp. (i.e., C. ubiquitum [25], C. suis [22], and C. canis [21]). At least two replicates were included in the PCR analysis of each target for each sample. To control for contamination and ensure validity during the PCR process, we used C. hominis from patients with diarrhea as a positive control. RNase-free water was used as a negative control.
Nucleotide sequencing and analysis
All amplified PCR products were visualized through 1% agarose gel electrophoresis (Fig. 2). All successful amplifications were then sent to Sangon Biotech (Shanghai) Co., Ltd. (Shanghai, China) for bidirectional sequencing. Subsequently, all nucleotide sequences were analyzed in MEGA version 11.0.13 [39] and compared with the NCBI GenBank database (http://www.ncbi.nlm.nih.gov/) to infer species/genotypes/subtypes of Cryptosporidium spp.
 |
Figure 2 Representative agarose gel image showing PCR amplification products of the small subunit (SSU) rRNA genes of Cryptosporidium spp. (expected band size 830 bp). Marker: DL 2000 DNA marker; N: negative control; P: positive control; Lanes 1–5: samples. |
Phylogenetic analysis
To determine the phylogeny of the Cryptosporidium spp. identified in this study, we constructed maximum likelihood (ML) trees based on the SSU rRNA gene of Cryptosporidium. DNA sequences downloaded from NCBI GenBank and those obtained in this study were aligned in MEGA to construct the ML tree. The best-fit nucleotide substitution models were tested with jModelTest version 2.1.4 [31]. Finally, ML trees were constructed with MEGA with a “GTR + I + G” substitution model for Cryptosporidium species, with 1,000 bootstrap replications.
Statistical analysis
The infection rates of Cryptosporidium spp. were determined according to the proportion of positive samples among the total rodent samples collected. The 95% confidence interval (CI) was used to quantify the precision of the positivity rate estimates. The chi-squared (χ2) test was used to statistically evaluate the variation in infection rates of Cryptosporidium spp. across rodent species, sexes, ages, sampling years, and sampling sites. All statistics were computed in R version 4.3.2 [32].
Results
Wild rodent community composition
A total of 273 wild rodents were captured from seven locations around Poyang Lake (Table 1). PCR and sequencing analysis of the COI gene identified two rodent species: A. agrarius (n = 125) and R. losea (n = 148) (Table 1). The rodent community was composed of four age groups: juvenile group (n = 17), sub-adult group (n = 89), adult group (n = 99), and old group (n = 68). The sex ratio of the community was balanced, with 142 males and 131 females (male-to-female ratio = 1.084, χ2 = 0.443, p = 0.506).
Frequency of Cryptosporidium spp.
Cryptosporidium spp. DNA was detected in 45 of 273 fecal samples, with an infection rate of 16.5% (95% CI: 12.3–21.4%). The infection rate of Cryptosporidium spp. was 20.3% (30/148, 95% CI: 13.9–26.7%) in A. agrarius and 12.0% (15/125, 95% CI: 6.9–19.0%) in R. losea. Although no significant differences were observed by host species (χ2 = 3.03, p = 0.082), sex (χ2 = 0.60, p = 0.439), or age group (χ2 = 1.12, p = 0.772), the infection rate was higher in A. agrarius than R. losea, and higher in male than female rodents. Additionally, the positivity rate increased with rodent age (Table 1).
Cryptosporidium species and genotypes
Seven known Cryptosporidium species and genotypes were identified through sequence and phylogenetic analyses of the SSU rRNA gene: rat genotype II (n = 13), C. apodemi (n = 12), C. muris (n = 6), C. suis (n = 5), C. ubiquitum (n = 5), rat genotype III (n = 4), and C. canis (n = 1) (Table 1, Fig. 3). The Cryptosporidium species and genotypes identified in this study exhibited 98.6%–100% similarity with the GenBank reference sequences and were clustered accordingly (Fig. 3). The subtyping of C. ubiquitum, C. suis, and C. canis failed.
 |
Figure 3 Phylogenetic analyses of the small subunit (SSU) rRNA genes of Cryptosporidium spp. with the maximum likelihood method (GTR + I + G model, 1000 replicates). Bootstrap values below 60% are not shown. Cryptosporidium species or genotypes identified in this study are highlighted with solid circles. Sequences are identified by their accession number, Cryptosporidium species, and host name. |
The distribution of Cryptosporidium spp. varied by rodent species, sex, age group, sampling year, and sampling site (Table 1). Analyses by rodent species indicated that rat genotype II was dominant in A. agrarius, whereas C. apodemi was dominant in R. losea. Analyses by sex indicated that rat genotype II and C. ubiquitum were dominant in male and female rodents, respectively. An assessment by rodent age indicated that C. muris and C. apodemi were dominant in the sub-adult group, whereas rat genotype II and rat genotype III were dominant in the adult and old groups, respectively. However, no dominant Cryptosporidium spp. was observed in the juvenile group, given the limited number of positive samples. An evaluation by sampling year indicated that C. apodemi, rat genotype III, and rat genotype II were most frequent in 2022, 2023, and 2024, respectively. An investigation by sampling site indicated that rat genotype II and C. apodemi were most frequent in YG; rat genotype III and C. apodemi were most frequent in LX; rat genotype II was most frequent in XJ and YX; and C. apodemi was most frequent in PY (Table 1).
Nucleotide sequence accession numbers
All Cryptosporidium spp. gene sequences identified in this study have been submitted to the NCBI GenBank database under accession numbers PV436589–PV436596.
Discussion
High infection rate of Cryptosporidium spp.
As important reservoirs for Cryptosporidium spp., wild rodents substantially contribute to pathogen transmission in the environment and to humans. Understanding the infection status, genetic characteristics, and zoonotic potential of Cryptosporidium spp. in wild rodents is crucial for preventing and controlling local cryptosporidiosis outbreaks, particularly in water-abundant ecosystems. This study provides the first evidence of a high Cryptosporidium spp. infection rate (16.5%, 45/273) in a wild rodent community in the Poyang Lake region. This infection rate is lower than the 20.5% (3848/18804) reported in wild rodents worldwide [47], yet higher than the 9.78% (345/3526) reported in China [51]. The Cryptosporidium spp. infection rate in rodents exhibits substantial geographical variation globally. The highest infection rates have been documented in Europe (28.0%, 1,860/6,638), followed by Asia (18.6%, 1,394/7,510), North America (15.2%, 1,265/8,299), Oceania (13.7%, 56/410), South America (7.3%, 11/150), and Africa (2.2%, 3/135) [47].
In China, Cryptosporidium infections in rodents have been reported across 18 of 34 provincial-level administrative divisions [51]. Regional distribution patterns have revealed notably high infection rates in southeastern districts, including Hainan Province (50.0%, 75/150) [50], Fujian (18.2%, 6/33) [27], and Guangdong (18.0%, 21/117) [4], as well as in our study area around Poyang Lake in Jiangxi province. In contrast, lower infection rates have been reported in the central and western regions, such as Chongqing (3.6%, 4/111) [4], Shanxi (3.8%, 2/53) [30], and Gansu (2.5%, 10/399) [45]. Although a nationwide systematic assessment of Cryptosporidium in wild rodents in China is lacking, available data indicate distinct geographical patterns. A recent review has shown generally higher prevalence in southeastern than northwestern China [51]. This southeast-to-northwest decreasing prevalence gradient mirrors the trend observed in human cryptosporidiosis [43], thereby suggesting elevated overall transmission risk in these southeastern regions. The high prevalence observed in the Poyang Lake region, which was consistent with this broader geographical trend, might be partly explained by its unique environmental conditions. The warm, humid climate and extensive wetlands in this region enhance oocyst survival and facilitate waterborne transmission [20]. These favorable conditions are further compounded by host-specific factors. Abundant food from surrounding agriculture supports high rodent densities, which in turn accelerate pathogen transmission [6]. Finally, strong anthropogenic influence is likely to be a considerable contributing factor. The Poyang Lake basin is characterized by a dense human population and large-scale livestock farming. Our detection of multiple zoonotic Cryptosporidium species in this context provides direct evidence of active transmission chains involving wildlife, livestock, and humans. This confluence of factors establishes the region as a “One Health” hotspot requiring close public health monitoring.
Of course, prevalence is a complex trait influenced not only by broad ecological conditions but also by host characteristics, such as species, sex, and age [12]. In this study, although no statistically significant differences in Cryptosporidium positivity rates were observed among rodent groups, numerically higher infection rates were found in A. agrarius (20.3%, 30/148) than R. losea (12.0%, 15/125), and in males (18.3%, 26/142) than females (14.5%, 19/131); moreover, a gradually increasing trend was observed with age (Table 1). These findings highlight the need for systematic, large-scale surveillance of Cryptosporidium spp. in wild rodents in this region, combined with detailed analysis of their zoonotic risks.
Zoonotic Cryptosporidium species
The need for surveillance is further underscored by our identification of four zoonotic species, C. ubiquitum, C. muris, C. canis, and C. suis, in these rodents, which might serve as reservoirs for human cryptosporidiosis. Among these, C. ubiquitum, which is frequently reported in humans [25], and is a geographically widespread species with a diverse range of mammalian hosts, including wild and domesticated ruminants, rodents, carnivores, and primates [14]. The role of rodents as an important reservoir for C. ubiquitum is highlighted by the identification of identical subtypes (XIIb and XIId) in rodents, humans, and the drinking water sources linking them, thus strongly suggesting a pathway for waterborne zoonotic transmission [25]. Our detection of C. ubiquitum in A. agrarius and R. losea – two dominant rodent species that frequently inhabit agricultural and residential areas – highlights a direct public health risk at the human-wildlife interface [6]. Similarly, the presence of C. muris in this study further reinforces wild rodents as important reservoirs. Although C. muris is known for its vast host range within rodent families, including Muridae, Cricetidae, Sciuridae, Caviidae, and Rhizomyidae [27], its importance as a human pathogen affecting primarily immunocompromised individuals in areas with poor sanitation cannot be overlooked [40, 47]. Despite being traditionally associated with domestic animals and humans [15], C. canis is increasingly being detected in wild canids [48, 41] and various small mammals [49]. Our detection of C. canis in A. agrarius and R. losea, in agreement with its discovery in other wild small mammals, adds substantial evidence indicating the sylvatic transmission cycle of C. canis. Given their ecological overlap with both wild and domestic canid populations, our findings suggest that these rodents might serve as potential bridge hosts, thus posing a transmission risk to domestic animals and humans. Cryptosporidium suis, a pathogen affecting primarily domestic pigs [34], has now been detected in diverse wildlife, including wild boar (Sus scrofa), red deer (Cervus elaphus), and red fox (Vulpes vulpes) [2, 7, 37]. Importantly, its presence in rodents is increasingly recognized, and it has been reported in species including the yellow-throated mouse (A. flavicollis) [7] and Brandt’s vole (Lasiopodomys brandtii) [16]. Our detection of C. suis in A. agrarius and R. losea further expands its potential rodent reservoir. However, the roles of rodents in the epidemiology of C. suis require careful interpretation. Distinguishing between mechanical carriage from a contaminated environment and a stable transmission cycle involving rodent-adapted subtypes is crucial for assessing zoonotic risk and warrants further investigation [12].
Our study also identified three dominant non-zoonotic species/genotypes in rodents: rat genotype II, rat genotype III, and C. apodemi. These species are largely considered specific to rodents: rat genotypes II and III infect primarily the house mouse (Mus musculus) [37], black rat (R. rattus) [18], and brown rat (R. norvegicus) [29], whereas C. apodemi is adapted to Apodemus spp. [5]. In this study, rat genotype II (n = 13) and C. apodemi (n = 12) were predominant in local wild rodents.
Implications for public health in lake ecosystems
In this study, the detection of zoonotic species/genotypes, such as C. suis, C. muris, C. canis, and C. ubiquitum, in the wild rodent communities suggests a risk of pathogen transmission from wildlife reservoirs to humans. Egan et al. [12] have demonstrated that synanthropic rodents have a high prevalence of Cryptosporidium infection and consequently are likely to contaminate source water and wastewater. Similarly, the common occurrence of Cryptosporidium rat genotype IV and C. muris suggested that rodents are important sources of enteric pathogens detected in wastewater samples in Guangzhou, China [13]. Moreover, the detection of C. muris and Cryptosporidium rat genotype IV in Laguna Lake in the Philippines has revealed rodent fecal pollution of a lake system [9]. In another study in South Australia, C. muris, C. ubiquitum, and C. tyzzeri were among the dominant species detected in source water [38]. The risk of rodent-associated Cryptosporidium species and genotypes commonly identified in source water and wastewater should be carefully estimated. In particular, the Poyang Lake region is a critical ecological landscape and primary water source for human consumption, crop irrigation, and livestock husbandry. Therefore, determining the diversity of waterborne protozoan parasites and the host-parasite pathogen spectrum is critical for understanding parasite evolution and informing the development of effective prevention and control strategies.
Unfortunately, we were unable to subtype Cryptosporidium spp. by amplifying the gp60 gene in the positive samples. For species with established gp60 subtyping tools (C. ubiquitum, C. canis, and C. suis), our amplification attempts were unsuccessful. No specific products were yielded for C. ubiquitum and C. canis, whereas for C. suis, sequencing of amplicons revealed non-specific bacterial DNA. These outcomes suggested that these samples contained an extremely low parasite load insufficient for amplification of the single-copy gp60 target gene. For other detected taxa, such as C. apodemi and the rat genotypes, standardized subtyping assays are not yet available. Future efforts should aim to clarify the zoonotic potential and transmission dynamics of Cryptosporidium spp. in this area. To overcome the issue of low-parasite-load samples, future studies might require more advanced methods, such as whole-genome amplification or targeted next-generation sequencing. Furthermore, a systematic investigation of a wider range of wild rodents should be conducted to more accurately estimate the prevalence and diversity of Cryptosporidium spp. in local wildlife.
Conclusions
This study provides, to our knowledge, the first assessment of Cryptosporidium spp. in wild rodents around Poyang Lake, China. We observed a 16.5% overall infection rate in the wild rodent community composed of A. agrarius and R. losea. Seven Cryptosporidium species/genotypes were identified, including four zoonotic species (C. muris, C. suis, C. ubiquitum, and C. canis) and three rodent-adapted species/genotypes (rat genotype II, rat genotype III, and C. apodemi). These findings highlight the roles of wild rodents as reservoirs of Cryptosporidium diversity. The presence of zoonotic Cryptosporidium poses a potential public health risk to the local environment. Given the ecological importance of Poyang Lake as a crucial freshwater resource, systematic surveillance of Cryptosporidium in both rodents and their environment is essential for assessing the zoonotic transmission risks.
Acknowledgments
We thank Dr. Chen Zhang at the Institute of Subtropical Agriculture, Chinese Academy of Sciences, for assistance in identifying rodent host species during this study.
Funding
This work was supported by the National Natural Science Foundation of China (No. 32160264 to X. Weng, No. 82372283 to Y. Shen), the NHC Key Laboratory of Echinococcosis Prevention and Control (No. 2024WZK1004 to Q. Zuo), and the Natural Science Foundation of Jiangxi Province (No. 20242BAB20266 to X. Weng).
Conflicts of interest
The authors have no commercial or other associations that represent a conflict of interest.
These authors contributed equally.
References
- Barbieri R, Signoli M, Chevé D, Costedoat C, Tzortzis S, Aboudharam G, Raoult D, Drancourt M. 2020. Yersinia pestis: the natural history of plague. Clinical Microbiology Reviews, 34, e00044–19. [Google Scholar]
- Barrera JP, Carmena D, Rodríguez E, Checa R, López AM, Fidalgo LE, Gálvez R, Marino V, Fuentes I, Miró G, Montoya A. 2020. The red fox (Vulpes vulpes) as a potential natural reservoir of human cryptosporidiosis by Cryptosporidium hominis in northwest Spain. Transboundary and Emerging Diseases, 67(5), 2172–2182. [Google Scholar]
- Chen JJ, Xu Q, Wang T, Meng FF, Li ZW, Fang LQ, Liu W. 2022. A dataset of diversity and distribution of rodents and shrews in China. Scientific Data, 9, 304. [CrossRef] [PubMed] [Google Scholar]
- Chen YW, Zheng WB, Zhang NZ, Gui BZ, Lv QY, Yan JQ, Zhao Q, Liu GH. 2019. Identification of Cryptosporidium viatorum XVa subtype family in two wild rat species in China. Parasites & Vectors, 12, 502. [CrossRef] [PubMed] [Google Scholar]
- Čondlová Š, Horčičková M, Sak B, Květoňová D, Hlásková L, Konečný R, Stanko M, McEvoy J, Kváč M. 2018. Cryptosporidium apodemi sp. n. and Cryptosporidium ditrichi sp. n. (Apicomplexa: cryptosporidiidae) in Apodemus spp. European Journal of Protistology, 53, 1–12. [Google Scholar]
- Dai N, Lu P, Zhang M, Zhang G, Zhang C, Pan Y. 2018. Small mammal communities on beaches and lakeside farmland in the Poyang Lake region after the Three-Gorges Project. Mammalia, 82, 438–448. [Google Scholar]
- Danišová O, Valenčáková A, Stanko M, Luptáková L, Hatalová E, Čanády A. 2017. Rodents as a reservoir of infection caused by multiple zoonotic species/genotypes of C. parvum, C. hominis, C. suis, C. scrofarum, and the first evidence of C. muskrat genotypes I and II of rodents in Europe. Acta Tropica, 172, 29–35. [CrossRef] [PubMed] [Google Scholar]
- Dashti A, Köster PC, Bailo B, De Las Matas AS, Habela MÁ, Rivero-Juarez A, Vicente J, Serrano E, Arnal MC, De Luco DF, Morrondo P, Armenteros JA, Balseiro A, Cardona GA, Martínez-Carrasco C, Ortiz JA, Carpio AJ, Calero-Bernal R, González-Barrio D, Carmena D. 2023. Occurrence and limited zoonotic potential of Cryptosporidium spp., Giardia duodenalis, and Balantioides coli infections in free-ranging and farmed wild ungulates in Spain. Research in Veterinary Science, 159, 189–197. [CrossRef] [PubMed] [Google Scholar]
- Dela Peña LBRO, Vejano MRA, Rivera WL. 2021. Molecular surveillance of Cryptosporidium spp. for microbial source tracking of fecal contamination in Laguna Lake, Philippines. Journal of Water and Health, 19, 534–544. [Google Scholar]
- Desai AN. 2020. Cryptosporidiosis. JAMA, 323(3), 288. [Google Scholar]
- Efstratiou A, Ongerth JE, Karanis P. 2017. Waterborne transmission of protozoan parasites: review of worldwide outbreaks–an update 2011–2016. Water Research, 114, 14–22. [CrossRef] [PubMed] [Google Scholar]
- Egan S, Barbosa AD, Feng Y, Xiao L, Ryan U. 2024. Critters and contamination: zoonotic protozoans in urban rodents and water quality. Water Research, 251, 121165. [CrossRef] [PubMed] [Google Scholar]
- Fan Y, Wang X, Yang R, Zhao W, Li N, Guo Y, Xiao L, Feng Y. 2021. Molecular characterization of the waterborne pathogens Cryptosporidium spp.; Giardia duodenalis, Enterocytozoon bieneusi, Cyclospora cayetanensis and Eimeria spp. in wastewater and sewage in Guangzhou, China. Parasites & Vectors, 14, 66. [CrossRef] [PubMed] [Google Scholar]
- Fayer R, Santín M, Macarisin D. 2010. Cryptosporidium ubiquitum n. sp. in animals and humans. Veterinary Parasitology, 172(1–2), 23–32. [Google Scholar]
- Fayer R, Trout JM, Xiao L, Morgan UM, Lai AA, Dubey JP. 2001. Cryptosporidium canis n. sp. from domestic dogs. Journal of Parasitology, 87, 1415–1422. [Google Scholar]
- Feng S, Chang H, Wang Y, Huang C, Han S, He H. 2020. Molecular characterization of Cryptosporidium spp. in Brandt’s Vole in China. Frontiers in Veterinary Science, 7, 300. [Google Scholar]
- Feng S-Y, Chang H, Luo J, Huang J-J, He H-X. 2019. First report of Enterocytozoon bieneusi and Cryptosporidium spp. in peafowl (Pavo cristatus) in China. International Journal for Parasitology: Parasites and Wildlife, 9, 1–6. [Google Scholar]
- García-Livia K, Fernández-Álvarez Á, Feliu C, Miquel J, Quilichini Y, Foronda P. 2022. Cryptosporidium spp. in wild murids (Rodentia) from Corsica, France. Parasitology Research, 121, 345–354. [CrossRef] [PubMed] [Google Scholar]
- Guo ZN, Su CH, Chen GW, Zheng ZM. 2014. Study on population age determination of Rattus losea. Chinese Journal of Hygienic Insecticides & Equipments, 20, 332–335. [Google Scholar]
- Harris J. 2016. Poyang Lake, Yangtze River Basin, China, in The wetland book: distribution, description and conservation, Finlayson MC, Davidson NC, Milton GR, Prentice C, Editors. Springer: Dordrecht. pp. 1–9. [Google Scholar]
- Jiang W, Roellig DM, Guo Y, Li N, Feng Y, Xiao L. 2021. Development of a subtyping tool for zoonotic pathogen Cryptosporidium canis. Journal of Clinical Microbiology, 59, e02474–20. [Google Scholar]
- Lebbad M, Grüttner J, Beser J, Lizana V, Dea-Ayuela MA, Oropeza-Moe M, Carmena D, Stensvold CR. 2024. Complete sequencing of the Cryptosporidium suis gp60 gene reveals a novel type of tandem repeats – implications for surveillance. Infection, Genetics and Evolution, 122, 105614. [Google Scholar]
- Li F, Zhang Z, Hu S, Zhao W, Zhao J, Kváč M, Guo Y, Li N, Feng Y, Xiao L. 2020. Common occurrence of divergent Cryptosporidium species and Cryptosporidium parvum subtypes in farmed bamboo rats (Rhizomys sinensis). Parasites & Vectors, 13, 149. [Google Scholar]
- Li M, Xu JL, Yin WS, Ao CH, Song ZS, Yang ZX. 2023. Variations of morphological characteristics and reproductive characteristics of Apodemus agrarius in Xifeng County. Chinese Agricultural Science Bulletin, 39, 150–156. [Google Scholar]
- Li N, Xiao L, Alderisio K, Elwin K, Cebelinski E, Chalmers R, Santin M, Fayer R, Kvac M, Ryan U, Sak B, Stanko M, Guo Y, Wang L, Zhang L, Cai J, Roellig D, Feng Y. 2014. Subtyping Cryptosporidium ubiquitum, a zoonotic pathogen emerging in humans. Emerging Infectious Diseases, 20, 217–224. [CrossRef] [PubMed] [Google Scholar]
- Li S, Zou Y, Wang P, Qu MR, Zheng WB, Wang P, Chen XQ, Zhu XQ. 2021. Prevalence and multilocus genotyping of Cryptosporidium spp. in cattle in Jiangxi Province, southeastern China. Parasitology Research, 120, 1281–1289. [CrossRef] [PubMed] [Google Scholar]
- Lv C, Zhang L, Wang R, Jian F, Zhang S, Ning C, Wang H, Feng C, Wang X, Ren X, Qi M, Xiao L. 2009. Cryptosporidium spp. in wild, laboratory, and pet rodents in china: prevalence and molecular characterization. Applied and Environmental Microbiology, 75, 7692–7699. [CrossRef] [PubMed] [Google Scholar]
- Němejc K, Sak B, Květoňová D, Hanzal V, Janiszewski P, Forejtek P, Rajský D, Ravaszová P, McEvoy J, Kváč M. 2013. Cryptosporidium suis and Cryptosporidium scrofarum in Eurasian wild boars (Sus scrofa) in central Europe. Veterinary Parasitology, 197, 504–508. [Google Scholar]
- Ng-Hublin JS, Singleton GR, Ryan U. 2013. Molecular characterization of Cryptosporidium spp. from wild rats and mice from rural communities in the Philippines. Infection, Genetics and Evolution, 16, 5–12. [Google Scholar]
- Ni HB, Sun YZ, Qin SY, Wang YC, Zhao Q, Sun ZY, Zhang M, Yang D, Feng ZH, Guan ZH, Qiu HY, Wang HX, Xue NY, Sun HT. 2021. Molecular detection of Cryptosporidium spp. and Enterocytozoon bieneusi infection in wild rodents from six provinces in China. Frontiers in Cellular and Infection Microbiology, 11, 783508. [CrossRef] [PubMed] [Google Scholar]
- Posada D, Crandall KA. 1998. MODELTEST: testing the model of DNA substitution. Bioinformatics, 14, 817–818. [CrossRef] [PubMed] [Google Scholar]
- R Core Team. 2020. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. [Google Scholar]
- Robins JH, Hingston M, Matisoo‐Smith E, Ross HA. 2007. Identifying Rattus species using mitochondrial DNA. Molecular Ecology Notes, 7, 717–729. [Google Scholar]
- Ryan UM, Monis P, Enemark HL, Sulaiman I, Samarasinghe B, Read C, Buddle R, Robertson I, Zhou L, Thompson RC, Xiao L. 2004. Cryptosporidium suis n. sp. (Apicomplexa: Cryptosporidiidae) in pigs (Sus scrofa). Journal of Parasitology, 90(4), 769–773. [CrossRef] [PubMed] [Google Scholar]
- Ryan UM, Feng Y, Fayer R, Xiao L. 2021. Taxonomy and molecular epidemiology of Cryptosporidium and Giardia – a 50-year perspective (1971–2021). International Journal for Parasitology, 51, 1099–1119. [CrossRef] [PubMed] [Google Scholar]
- Schönrich G, Rang A, Lütteke N, Raftery MJ, Charbonnel N, Ulrich RG. 2008. Hantavirus‐induced immunity in rodent reservoirs and humans. Immunological Reviews, 225, 163–189. [Google Scholar]
- Silva SO, Richtzenhain LJ, Barros IN, Gomes AM, Silva AV, Kozerski ND, de Araújo Ceranto JB, Keid LB, Soares RM. 2013. A new set of primers directed to 18S rRNA gene for molecular identification of Cryptosporidium spp. and their performance in the detection and differentiation of oocysts shed by synanthropic rodents. Experimental Parasitology, 135, 551–557. [Google Scholar]
- Swaffer B, Abbott H, King B, van der Linden L, Monis P. 2018. Understanding human infectious Cryptosporidium risk in drinking water supply catchments. Water Research, 138, 282–292. [Google Scholar]
- Tamura K, Stecher G, Kumar S. 2021. MEGA11: Molecular evolutionary genetics analysis version 11. Molecular Biology and Evolution, 38, 3022–3027. [CrossRef] [PubMed] [Google Scholar]
- Tiangtip R, Jongwutiwes S. 2002. Molecular analysis of Cryptosporidium species isolated from HIV‐infected patients in Thailand. Tropical Medicine & International Health, 7, 357–364. [Google Scholar]
- Trout JM, Santín M, Fayer R. 2006. Giardia and Cryptosporidium species and genotypes in coyotes (Canis latrans). Journal of Zoo and Wildlife Medicine, 37, 141–144. [Google Scholar]
- Wang P, Li S, Zou Y, Du Z-C, Song DP, Wang P, Chen XQ. 2022. The infection and molecular characterization of Cryptosporidium spp. in diarrheic pigs in southern China. Microbial Pathogenesis, 165, 105459. [CrossRef] [PubMed] [Google Scholar]
- Wang X, Shen YJ, Cao JP. 2022. Epidemic status and prevention and control progress of cryptosporidiosis in China. Journal of Tropical Diseases and Parasitology, 20, 136–148. [Google Scholar]
- Xiao L, Singh A, Limor J, Graczyk TK, Gradus S, Lal A. 2001. Molecular characterization of Cryptosporidium oocysts in samples of raw surface water and wastewater. Applied and Environmental Microbiology, 67, 1097–1101. [Google Scholar]
- Xu J, Liu H, Jiang Y, Jing H, Cao J, Yin J, Li T, Sun Y, Shen Y, Wang X. 2022. Genotyping and subtyping of Cryptosporidium spp. and Giardia duodenalis isolates from two wild rodent species in Gansu Province, China. Scientific Reports, 12, 12178. [CrossRef] [PubMed] [Google Scholar]
- Yu Z, Wen X, Huang X, Yang R, Guo Y, Feng Y, Xiao L, Li N. 2020. Molecular characterization and zoonotic potential of Enterocytozoon bieneusi, Giardia duodenalis and Cryptosporidium sp. in farmed masked palm civets (Paguma larvata) in southern China. Parasites & Vectors, 13, 403. [CrossRef] [PubMed] [Google Scholar]
- Zhang K, Fu Y, Li J, Zhang L. 2022. Public health and ecological significance of rodents in Cryptosporidium infections. One Health, 14, 100364. [CrossRef] [PubMed] [Google Scholar]
- Zhang S, Tao W, Liu C, Jiang Y, Wan Q, Li Q, Yang H, Lin Y, Li W. 2016. First report of Cryptosporidium canis in foxes (Vulpes vulpes) and raccoon dogs (Nyctereutes procyonoides) and identification of several novel subtype families for Cryptosporidium mink genotype in minks (Mustela vison) in China. Infection, Genetics and Evolution, 41, 21–25. [Google Scholar]
- Zhang X, Jian Y, Li X, Ma L, Karanis G, Karanis P. 2018. The first report of Cryptosporidium spp. in Microtus fuscus (Qinghai vole) and Ochotona curzoniae (wild plateau pika) in the Qinghai-Tibetan Plateau area, China. Parasitology Research, 117, 1401–1407. [CrossRef] [PubMed] [Google Scholar]
- Zhao W, Zhou H, Huang Y, Xu L, Rao L, Wang S, Wang W, Yi Y, Zhou X, Wu Y, Ma T, Wang G, Hu X, Peng R, Yin F, Lu G. 2019. Cryptosporidium spp. in wild rats (Rattus spp.) from the Hainan Province, China: Molecular detection, species/genotype identification and implications for public health. International Journal for Parasitology: Parasites and Wildlife, 9, 317–321. [CrossRef] [Google Scholar]
- Zhou SS, Jiang YY, Cao JP. 2024. Cryptosporidium infection in rodents in China. Chinese Journal of Parasitology and Parasitic Diseases, 42, 512–520. [Google Scholar]
Cite this article as: Zuo Q, Weng X, Wang X, Liu H, Di M, Zhang X, Zhou B, Xue C, Lu P, Peng X, Shen Y & Cao J. 2025. Cryptosporidium spp. in wild rodents in the Poyang Lake region, China. Parasite 32, 63. https://doi.org/10.1051/parasite/2025056.
All Tables
All Figures
 |
Figure 1 Study area and sampling sites (marked with yellow squares) around Poyang Lake, Jiangxi Province, China. |
| In the text | |
 |
Figure 2 Representative agarose gel image showing PCR amplification products of the small subunit (SSU) rRNA genes of Cryptosporidium spp. (expected band size 830 bp). Marker: DL 2000 DNA marker; N: negative control; P: positive control; Lanes 1–5: samples. |
| In the text | |
 |
Figure 3 Phylogenetic analyses of the small subunit (SSU) rRNA genes of Cryptosporidium spp. with the maximum likelihood method (GTR + I + G model, 1000 replicates). Bootstrap values below 60% are not shown. Cryptosporidium species or genotypes identified in this study are highlighted with solid circles. Sequences are identified by their accession number, Cryptosporidium species, and host name. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


