Figure 1
Download original image
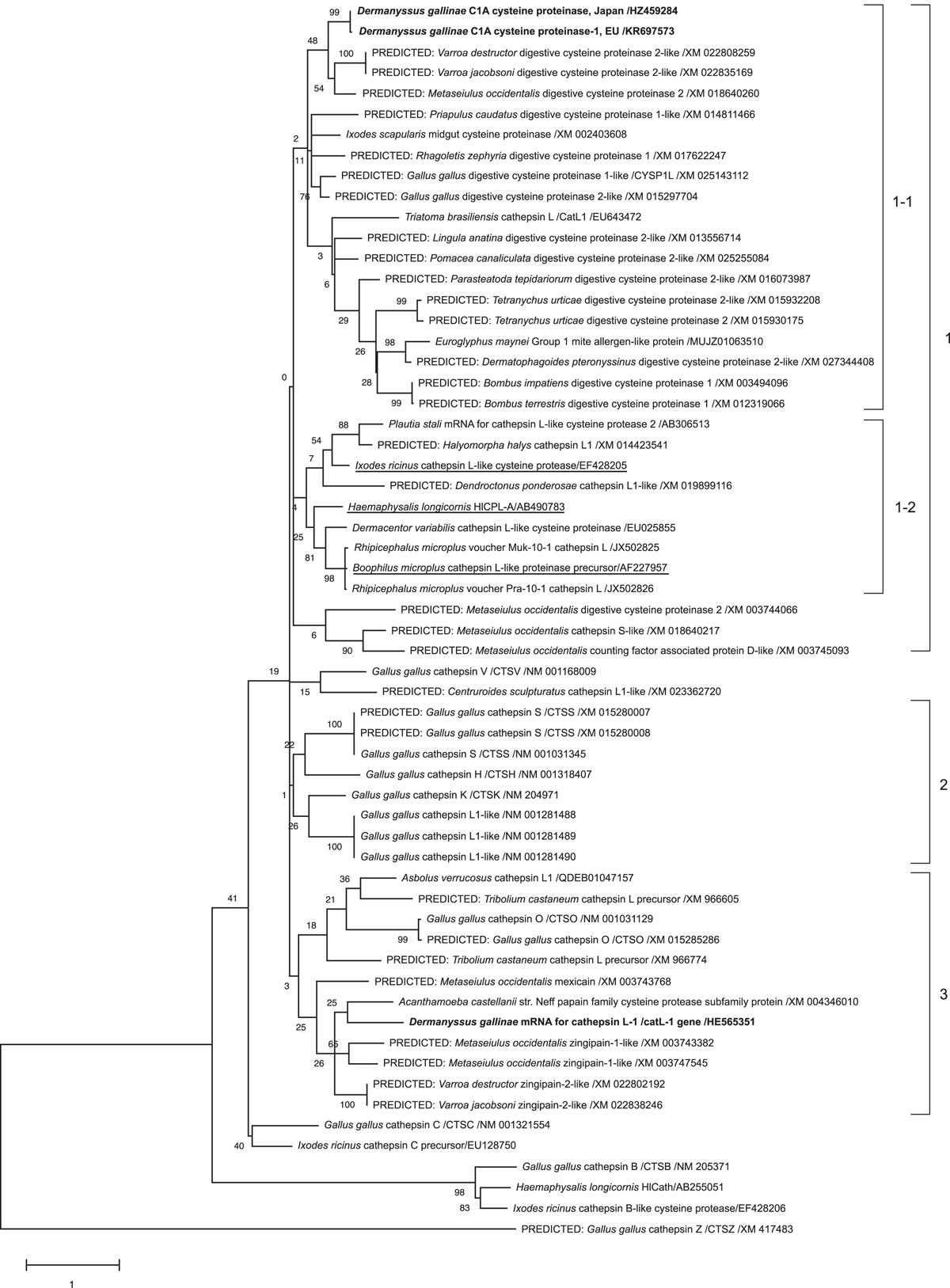
Phylogenetic tree based on the nucleotide sequences of the open reading frames of the cysteine protease genes in poultry red mites (PRMs, Dermanyssus gallinae), arthropods, including other mites and ticks, chickens, and other species such as insects and invertebrates. The tree was constructed using the maximum-likelihood method with MEGA X software [14]. The numbers on the right indicate the clusters. Cluster 1: This cluster was divided into two sub-clusters. The digestive cysteine proteases from various species were classified into cluster 1–1, and cysteine proteases from PRMs (Deg-CPR-1) belonged to this cluster (bold font). Sub-cluster 1–2 included cathepsin L-like proteases and some cysteine proteases that are proposed to play a role in haemoglobin digestion in ticks (underlined). Cluster 2: This cluster comprised the cysteine proteases and cathepsins S, H, K, and L of chickens. Cluster 3: The cysteine proteases from mites, different from those in cluster 1 were included in this cluster, and another cysteine protease from PRMs, Dg-CatL-1, was also included (bold font).
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


