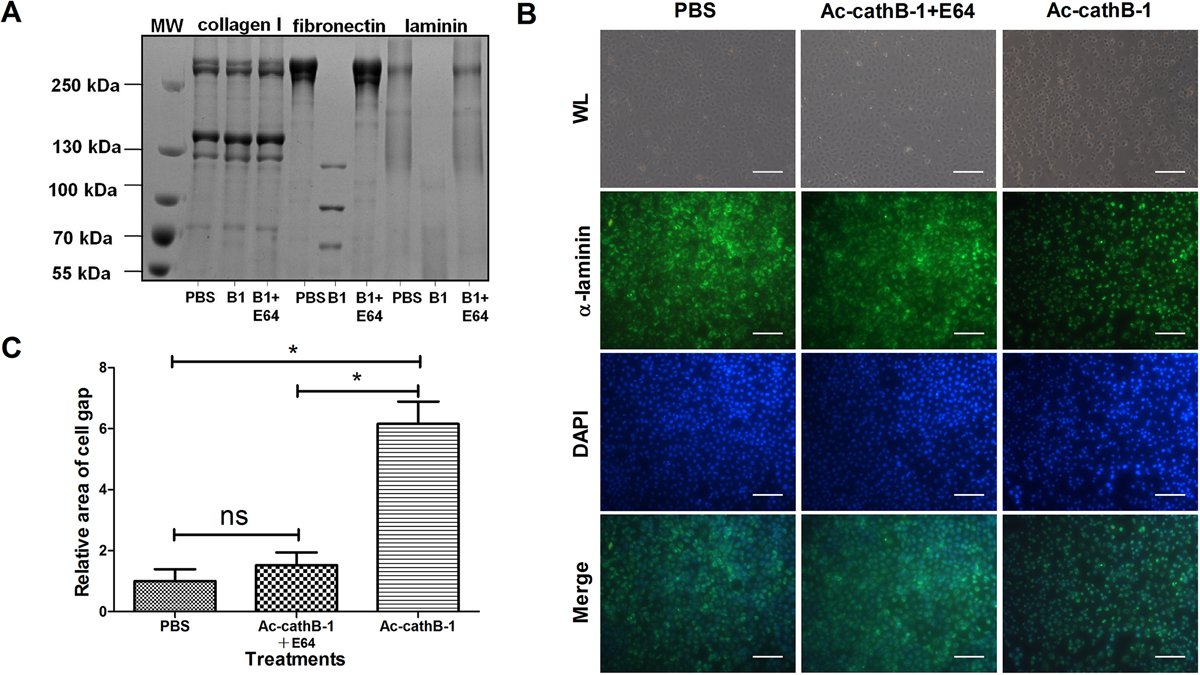
Figure 4.
Download original image
Assessment of the hydrolytic activity of activated rAc-cathB-1. (A) Activated rAc-cathB-1 completely degraded fibronectin and laminin but did not cleave type I collagen over a 12-h period. In the incubation buffer (pH 6.0), 10 μg of the respective substrates was treated with equal volumes of PBS, activated rAc-cathB-1, or activated rAc-cathB-1 plus E64 at 37 °C for 12 h. After incubation, all samples were analyzed on a 5% SDS-PAGE gel followed by Coomassie brilliant blue staining. Molecular weight markers (in kDa) are listed on the side. The substrate used for each experiment is indicated at the top of the gel, and the presence or absence of a recombinant protease or protease inhibitor is indicated at the bottom of each lane. (B) Influence of rAc-cathB-1 on IEC-6 monolayer. IEC-6 cells were grown to confluence and equal amounts of rAc-cathB-1 with or without E64 were applied to cells for 2 h. The blank was made of IEC-6 with PBS added. After incubation, the adherent IEC-6 partly rounded up and the integrity of the cell sheet was disrupted by activated rAc-cathB-1. The cytoplasm and ECM were then labeled with an anti-laminin antibody (green), and the nucleus was stained with DAPI. On the merged images, the dark regions represent the intercellular space. (C) Statistical analysis of the changes to the intercellular space of IEC-6 cells. The dark area was measured and analyzed. The difference between the means for each group of samples was estimated using one-way ANOVA followed by Duncan’s multiple comparison test. Asterisk (*), P < 0.05; WL, white light; ns: not significant; and bar = 100 μm.
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


